User:IsadoraofIbiza/sandbox/Chloroplast

the many-fruited thyme moss
Chloroplasts /ˈklɔːrəplæsts/ are organelles found in plant cells and some other eukaryotic organisms. As well as conducting photosynthesis, they carry out almost all fatty acid synthesis in plants, and are involved in a plant's immune response. A chloroplast is a type of plastid which specializes in photosynthesis. During photosynthesis, chloroplasts capture the sun's light energy, and store it in the energy storage molecules ATP and NADPH while freeing oxygen from water. They then use the ATP and NADPH to make organic molecules from carbon dioxide in a process known as the Calvin cycle.[1]
The word chloroplast (χλωροπλάστης) is derived from the Greek words chloros (χλωρός), which means green, and plastes (πλάστης), which means "the one who forms".[2]
Chloroplast lineages and evolution
[edit]Chloroplasts are one of the many different types of organelles in the plant cell. However, unlike most cellular organelles, they are considered to have originated from cyanobacteria through endosymbiosis—when a eukaryotic cell engulfed a photosynthesizing cyanobacterium which remained and became a permanent resident in the cell. Mitochondria are thought to have come from a similar event, where a ærobic prokaryote was engulfed.[3] This origin of chloroplasts was first suggested by Konstantin Mereschkowski in 1905[4] after Andreas Schimper observed that chloroplasts closely resemble cyanobacteria in 1883.[5]
Chloroplasts are similar to mitochondria in that they both originate from an endosymbiotic event, but chloroplasts are found only in plants and some protists.
Cyanobacterial ancestor
[edit]Cyanobacteria are considered the ancestors of chloroplasts. They are sometimes called blue-green algae even though they are prokaryotes. They are a diverse phylum of bacteria capable of carrying out photosynthesis, and are gram-negative, meaning they have two cell membranes. They also contain a thick peptidoglycan cell wall which is located between their two cell membranes.[6] Like chloroplasts, they have thylakoids inside of them.
Primary endosymbiosis
[edit]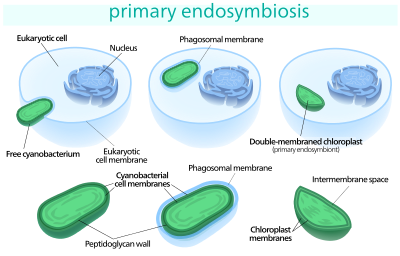
Somewhere around a billion years ago,[7] a free-living cyanobacterium entered an early eukaryotic cell, either as food or an internal parasite,[3] and managed to escape the phagocytic vacuole it was contained in.[8] The two innermost lipid-bilayer membranes that surround all chloroplasts[9] correspond to the outer and inner membranes of the ancestral cyanobacterium's gram negative cell wall,[10][11][12] and not the phagosomal membrane from the host, which was probably lost.[12] The new cellular resident quickly became an advantage, providing food for the eukaryotic host, which allowed it to live within it.[3] Over time, the cyanobacterium was assimilated, and many of its genes were lost or transferred to the nucleus of the host.[13] Some of its proteins were then synthesized in the cytoplasm of the host cell, and imported back into the chloroplast.[14][13]
According to the serial endosymbiosis theory, chloroplasts are believed to have arisen after mitochondria, since all eukaryotes contain mitochondria, but not all have chloroplasts.[3][15]
Whether or not chloroplasts came from a single endosymbiotic event, or many independent engulfments across various eukaryotic lineages has been long debated, but it is now generally held that all organisms with chloroplasts either share a single ancestor or obtained their chloroplast from organisms that share a common ancestor that took in a cyanobacterium 600–1600 million years ago.[7]
| Glaucophyta | Chloroplast lineages A primary endosymbiosis event gave rise to three main lineages of chloroplasts in the glaucophytes, chlorophyta, and rhodophyta.[16] Some of these algae were subsequently engulfed by other algae, becoming secondary (or tertiary) endosymbionts.[8] a The apicomplexans (malaria parasites), contain a red algal endosymbiont with a non- photosynthetic chloroplast.[17] b 2–3 chloroplast membranes[8] a c 2–4 chloroplast membranes[8] d 3 chloroplast membranes[8] | |||||
| Chloroplastida |
Euglenophyta | |||||
| Chlorarachniophyta | ||||||
| Green algal dinophytes | ||||||
| Rhodophyceæ (Red algae) |
Apicomplexa a | |||||
| Peridinin-type dinophytes b | ||||||
| Cryptophyta | ||||||
| Haptophyta | Haptophyte dinophytes c | |||||
| Heterokontophyta | Diatom dinophytes d | |||||
| Primary endosymbiosis | Secondary endosymbiosis | Tertiary endosymbiosis | ||||
These chloroplasts, which can be traced back directly to a cyanobacterial ancestor are known as primary plastids.[18] All primary chloroplasts belong to one of three chloroplast lineages—the glaucophyte chloroplast lineage, the rhodophyte, or red algal chloroplast lineage, or the chloroplastidan, or green chloroplast lineage.[16] The second two are the largest,[12] and the green chloroplast lineage is the one that contains the land plants.[12]
Glaucophyta
[edit]The alga Cyanophora, a glaucophyte, is thought to be one of the first organisms to contain a chloroplast.[14] The glaucophyte chloroplast group is the smallest of the three primary chloroplast lineages, being found in only thirteen species,[12] and is thought to be the one that branched off the earliest.[12][7][19] Glaucophytes have chloroplasts which retain a peptidoglycan wall between their double membranes,[18] like their cyanobacterial parent.[6] For this reason, glaucophyte chloroplasts are also known as muroplasts.[18] Glaucophyte chloroplasts also contain concentric unstacked thylakoids which surround a carboxysome, an icosahedral structure that glaucophyte chloroplasts and cyanobacteria keep their carbon fixation enzyme rubisco in. The starch they synthesize collects outside the chloroplast.[8] Like cyanobacteria, glaucophyte chloroplast thylakoids are studded with light collecting structures called phycobilisomes.[8][18] For these reasons, glaucophyte chloroplasts are considered a primitive intermediate between cyanobacteria and the more evolved chloroplasts in red algae and plants.[18]
Rhodophyceæ (red algae)
[edit]The rhodophyte, or red algal chloroplast group is another large and diverse chloroplast lineage.[12] Rhodophyte chloroplasts are also called rhodoplasts,[18] literally "red chloroplasts".[21]
Rhodoplasts have a double membrane with an intermembrane space and unstacked thylakoids which contain phycobilisomes. Some contain pyrenoids.[18] Rhodoplasts have chlorophyll a and phycobilins[19] for photosynthetic pigments; the phycobillin phycoerytherin is responsible for giving many red algae their distinctive red color.[20] However, since they also contain the blue-green chlorophyll a and other pigments, many are reddish to purple from the combination.[18] The red phycoerytherin pigment is an adaptation to help red algae catch more sunlight in deep water[18]—as such, some red algae that live in shallow water have less phycoerytherin in their rhodoplasts, and can appear more greenish.[20] Rhodoplasts synthesize a form of starch called floridean,[18] which collects into granules outside the rhodoplast, in the cytoplasm of the red alga.[8]
Chloroplastida (green algae and plants)
[edit]The chloroplastidan chloroplasts, or green chloroplasts, are a large, highly diverse second primary chloroplast lineage. They are commonly known as the green algae and land plants.[22] They differ from glaucophyte and red algal chloroplasts in that they have lost their phycobilisomes, and contain chlorophyll b instead.[8] Most green chloroplasts are (obviously) green, though some, like some forms of Hæmatococcus pluvialis aren't, due to accessory pigments that override the chlorophylls' green colors. Chloroplastidan chloroplasts have lost the peptidoglycan wall between their double membrane, and have replaced it with an intermembrane space.[8]
Green algae and plants keep their starch inside their chloroplasts,[19][22][8] and in plants and some algae, the chloroplast thylakoids are arranged in grana stacks. Some green algal chloroplasts contain a structure called a pyrenoid,[8] which is functionally similar to the glaucophyte carboxysome in that it's where rubisco and CO2 is concentrated in the chloroplast.[23]

Secondary and tertiary endosymbiosis
[edit]Many other organisms obtained chloroplasts from the primary chloroplast lineages through secondary endosymbiosis—engulfing a red or green alga that contained a chloroplast. These chloroplasts are known as secondary plastids.[18]
While primary chloroplasts have a double membrane from their cyanobacterial ancestor, secondary chloroplasts have additional membranes outside of the original two, as a result of the secondary endosymbiotic event, when a nonphotosynthetic eukaryote engulfed an chloroplast-containing alga but failed to digest it—much like the cyanobacterium at the beginning of this story.[12] The engulfed alga was broken down, leaving only its chloroplast, and sometimes its cell membrane and nucleus, forming a chloroplast with three to four membranes[24]—the two cyanobacterial membranes, sometimes the eaten alga's cell membrane, and the phagosomal vacuole from the host's cell membrane.


The genes in the phagocytosed eukaryote's nucleus are often transferred to the secondary host's nucleus.[12] Cryptomonads and chlorarachniophytes retain the phagocytosed eukaryote's nucleus, an object called a nucleomorph,[12] located between the second and third membranes of the chloroplast.[14][8]
All secondary chloroplasts come from green and red algae—no secondary chloroplasts from glaucophytes have been observed, probably because glaucophytes are relatively rare in nature, making them less likely to have been taken up by another eukaryote.[12]
Green algal derived chloroplasts
[edit]Green algae have been taken up by the euglenids, chlorarachniophytes, and a group of dinoflagellates.[19] Many green algal derived chloroplasts contain pyrenoids, but unlike chloroplasts in their green algal ancestors, starch collects in granules outside the chloroplast.[8]
Euglenophytes
[edit]Euglenophytes are a group of common flagellated protists that contain chloroplasts derived from a green alga.[12] Euglenophyte chloroplasts have three membranes—it's thought that the membrane of the primary endosymbiont was lost, leaving the cyanobacterial membranes, and the secondary host's phagosomal membrane.[12] Euglenophyte chloroplasts have a pyrenoid and thylakoids stacked in groups of three. Starch is stored in the form of paramylon, which is contained in membrane-bound granules in the cytoplasm of the euglenophyte.[8][19]

Chlorarachniophytes
[edit]Chlorarachniophytes (/ˌklɔːrəˈrækni[invalid input: 'ɵ']ˌfaɪts/) are a rare group of organisms that also contain chloroplasts derived from green algae,[12] though their story is more complicated than that of the euglenophytes. The ancestor of chlorarachniophytes is thought to have been a chromalveolate, a eukaryote with a red algal derived chloroplast. It's then thought to have lost its first red algal chloroplast, and later engulfed a green alga, giving it its second, green algal derived chloroplast.[19]
Chlorarachniophyte chloroplasts are bounded by four membranes, except near the cell membrane, where the chloroplast membranes fuse into a double membrane.[8] Their thylakoids are arranged in loose stacks of three.[8] Chlorarachniophytes have a form of starch called chrysolaminarin, which they store in the cytoplasm,[19] often collected around the chloroplast pyrenoid, which bulges into the cytoplasm.[8]
Chlorarachniophyte chloroplasts are notable because the green alga they are derived from has not been completely broken down—its nucleus still persists as a nucleomorph[12] found between the second and third chloroplast membranes[8]—the periplastid space, which corresponds to the green alga's cytoplasm.[19]
Red algal derived chloroplasts
[edit]Like green algae, red algae have also been taken up in secondary endosymbiosis, though it's thought that all red algal derived chloroplasts are descended from a single red alga that was engulfed by an early chromalveolate, giving rise to the chromalveolates, some of which subsequently lost the chloroplast.[12][19][20] This is still debated though.[19][20]
Pyrenoids and stacked thylakoids are common in chromalveolate chloroplasts, and the outermost membrane of many are continuous with the rough endoplasmic reticulum and studded with ribosomes.[19][8] They have lost their phycobilisomes and exchanged them for chlorophyll c, which isn't found in primary red algal chloroplasts themselves.[8]
Cryptophytes
[edit]
Cryptophytes, or cryptomonads are a group of algae that contain a red-algal derived chloroplast. Cryptophyte chloroplasts contain a nucleomorph that superficially resembles that of the chlorarachniophytes.[12] Cryptophyte chloroplasts have four membranes, the outermost of which is continuous with the rough endoplasmic reticulum. They synthesize ordinary starch, which is stored in granules found in the periplastid space—outside the original double membrane, in the place that corresponds to the red alga's cytoplasm. Inside cryptophyte chloroplasts is a pyrenoid and thylakoids in stacks of two.[8]

Haptophytes
[edit]Haptophytes are similar and closely related to cryptophytes, and are thought to be the first chromalveolates to branch off.[19] but their chloroplasts lack a nucleomorph,[8][12] their thylakoids are in stacks of three, and they synthesize chrysolaminarin sugar, which they store completely outside of the chloroplast, in the cytoplasm of the haptophyte.[8]
Heterokontophytes
[edit]
The heterokontophytes, also known as the stramenopiles, are a very large and diverse group of algae that also contain red algal derived chloroplasts.[19] Heterokonts include the diatoms and the brown, golden,[20] and yellow-green algae. Heterokont chloroplasts are very similar to haptophyte chloroplasts, containing a pyrenoid, triplet thylakoids, and having an epiplastid membrane connected to the endoplasmic reticulum. Like haptophytes, heterokontophytes store sugar in chrysolaminarin granules in the cytoplasm.[8]
Heterokontophyte chloroplasts contain chlorophylls a and c,[12] but also have carotenoids which give them their many colors.[20]
In some cases, such secondary endosymbionts may have themselves been engulfed by still other eukaryotes,[25] thus forming tertiary endosymbionts.
Chromatophores
[edit]While most chloroplasts originate from that first set of endosymbiotic events, Paulinella chromatophora is an exception, which has acquired a photosynthetic cyanobacterial endosymbiont more recently. It is not closely related to chloroplasts of other eukaryotes.[26] Being in the early stages of endosymbiosis, Paulinella chromatophora can offer some insights into how chloroplasts evolved.[13][27] Paulinella cells contain one or two sausage shaped blue-green photosynthesizing structures called chromatophores,[13][27] descended from the cyanobacterium Synechococcus. Chromatophores cannot survive outside their host.[13] Chromatophore DNA is about a million base pairs long, containing around 850 protein encoding genes—far less than the three million base pair Synechococcus genome,[13] but much larger than the ~150,000 base pair genome of the more assimilated chloroplast.[28][29][30] Chromatophores have transferred much less of their DNA to the nucleus of their host. About 0.3–0.8% of the nuclear DNA in Paulinella is from the chromatophore, compared with 11–14% from the chloroplast in plants.[27]
Kleptoplastidy
[edit]In some groups of mixotrophic protists, like some dinoflagellates, chloroplasts are separated from a captured alga or diatom and used temporarily. These klepto chloroplasts may only have a lifetime of a few days and are then replaced.[31]
Chloroplast DNA
[edit]Chloroplasts have their own DNA,[32] often abbreviated as ctDNA,[33] or cpDNA.[34] It is also known as the plastome. Its existence was first proved in 1962,[28] and first sequenced in 1986—when two Japanese research teams sequenced the chloroplast DNA of liverwort and tobacco.[35] Since then, hundreds of chloroplast DNAs from various species have been sequenced—mostly those of land plants and green algae.[36]
Molecular structure
[edit]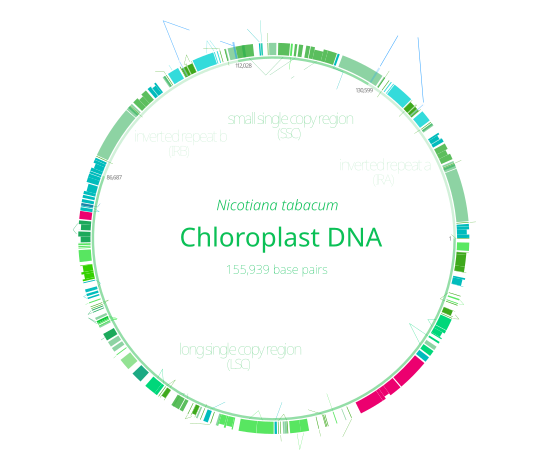
Chloroplast DNAs are circular, and are typically 120,000–170,000 base pairs long.[28][29][30] They can have a contour length of around 30–60 micrometers, and have a mass of about 80–130 million daltons.[37]
Most plants have their entire chloroplast genome combined into a single large ring, though dinoflagellate algae are a notable exception—their chloroplasts contain around forty small plasmids which each consist of one to three genes.[14][36] Many chloroplast DNAs contain two large inverted repeats about 20,000–25,000 base pairs long each,[30][38] which separate a long single copy section (LSC) from a short single copy section (SSC).[30] The inverted repeat regions are highly conserved among various plants, and accumulate few mutations.[30][38] Similar inverted repeats exist in the genomes of cyanobacteria, suggesting that they predate the chloroplast,[36] though some plants like peas have since lost the inverted repeats.[38][39] It is possible that the inverted repeats help stabilize the rest of the chloroplast genome, as chloroplast DNAs which have lost some of the inverted repeat segments tend to get rearranged more.[39] Chloroplast genomes as a whole are also generally well conserved among land plants.[30]
New chloroplasts may contain up to 100 copies of their DNA,[28] though the number of chloroplast DNA copies decreases to about 15–20 as the chloroplasts age.[40] They are usually packed into nucleoids which can contain several identical chloroplast DNA rings. Many nucleoids can be found in each chloroplast.[37]
Though chloroplast DNA is not associated with true histones,[3] in red algae, a histone-like chloroplast protein (HC) coded by the chloroplast DNA that tightly packs each chloroplast DNA ring into a nucleoid has been found.[41]
In primitive red algae, the chloroplast DNA nucleoids are clustered in the center of the chloroplast, while in green plants and green algae, the nucleoids are dispersed throughout the stroma.[41]
Genes
[edit]The chloroplast genome includes around 100 genes[14][29] which code for a variety of things, mostly to do with the protein pipeline and photosynthesis. As in prokaryotes, genes in chloroplast DNA are organized into operons.[14]
Among land plants, the contents of the chloroplast genome are fairly similar—they code for four ribosomal RNAs, 30–31 tRNAs, 21 ribosomal proteins, and four RNA polymerase subunits,[42][43] involved in protein synthesis. For photosynthesis, the chloroplast DNA includes genes for 28 thylakoid proteins and the large Rubisco subunit.[42] In addition, its genes encode eleven subunits of a protein complex which mediates redox reactions to recycle electrons,[44] which is similar to the NADH dehydrogenase found in mitochondria.[42][45]
Chloroplast genome reduction and protein synthesis
[edit]The chloroplast genome is considerably reduced compared to that of free-living cyanobacteria, but the parts that are still present show clear similarities with the cyanobacterial genome. Plastids may contain 60–100 genes whereas cyanobacteria often contain more than 1500 genes.[46] Over time, many parts of the chloroplast genome were transferred to the nuclear genome of the host,[28][29][47] a process called endosymbiotic gene transfer. In land plants, some 11–14% of the DNA in their nuclei can be traced back to the chloroplast.[27] There have been a few recent transfers of genes from the chloroplast DNA to the nuclear genome in land plants.[29]
Of the approximately three-thousand proteins found in chloroplasts, some 95% of them are encoded by nuclear genes. Many of the chloroplast's protein complexes consist of subunits from both the chloroplast genome and the host's nuclear genome. As a result, protein synthesis must be coordinated between the chloroplast and the nucleus. The chloroplast is mostly under nuclear control, though chloroplasts can also give out signals regulating gene expression in the nucleus.[48]
Protein synthesis within chloroplasts relies on an RNA polymerase coded by the chloroplast's own genome, which is related to RNA polymerases found in bacteria. Chloroplasts also contain a mysterious second RNA polymerase that is encoded by the plant's nuclear genome. The two RNA polymerases may recognize and bind to different kinds of promoters within the chloroplast genome.[49] The ribosomes in chloroplasts are similar to bacterial ribosomes.[42]
This section needs expansion with: Genome size differences between algae and land plants, chloroplast stuff coded by the nucleus, DNA replication, NADPH redox, special tRNA synthetases, etc.. You can help by adding to it. (January 2013) |
Structure
[edit]
In land plants, chloroplasts are generally lens-shaped, 5–8 μm in diameter and 1–3 μm thick.[50] Greater diversity in chloroplast shapes exists among the algae, which can have chloroplasts shaped like a net (e.g, Oedogonium),[51] a cup (e.g, Chlamydomonas),[52] a ribbon-like spiral around the edges of the cell (e.g, Spirogyra),[53] or slightly twisted bands at the cell edges (e.g, Sirogonium).[54] Some algae have two chloroplasts in each cell; they are star-shaped in Zygnema,[55] or may follow the shape of half the cell in order Desmidiales.[56] In some red algae, the chloroplast takes up most of the cell, with pockets for the nucleus and other organelles.[8]
All chloroplasts have at least three membrane systems—the outer chloroplast membrane, the inner chloroplast membrane, and the thylakoid system. Chloroplasts that are the product of secondary endosymbiosis may have additional membranes surrounding these three.[24] Inside the outer and inner chloroplast membranes is the chloroplast stroma, a semi-gel-like fluid[18] that makes up much of a chloroplast's volume, and in which the thylakoid system floats.
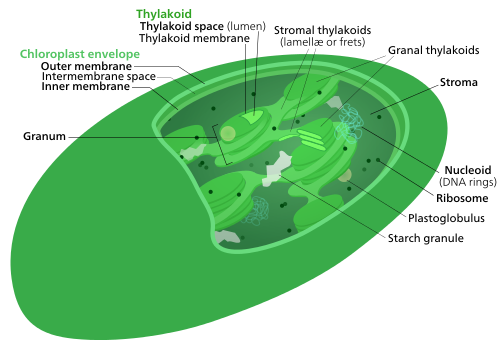
There are some common misconceptions about the outer and inner chloroplast membranes. The fact that chloroplasts are surrounded by a double membrane is often cited as evidence that they are the descendants of endosymbiotic cyanobacteria. This is often interpreted as meaning the outer chloroplast membrane is the product of the host's cell membrane infolding to form a vesicle to surround the ancestral cyanobacterium—which is not true—both chloroplast membranes are homologous to the cyanobacterium's original double membranes.[12]
The chloroplast double membrane is also often compared to the mitochondrial double membrane. This is not a valid comparison—the inner mitochondria membrane is used to run proton pumps and carry out oxidative phosphorylation across to generate ATP energy. The only chloroplast structure that can considered analogous to it is the internal thylakoid system. Even so, in terms of "in-out", the direction of chloroplast H+ ion flow is in the opposite direction compared to oxidative phosphorylation in mitochondria.[18][57] In addition, in terms of function, the inner chloroplast membrane, which regulates metabolite passage and synthesizes some materials, has no counterpart in the mitochondrion.[18]
Outer chloroplast membrane
[edit]The outer chloroplast membrane is a semi-porous membrane that small molecules and ions can easily diffuse across.[58] However, it's not permeable to larger proteins, so chloroplast polypeptides being synthesized in the cell cytoplasm must be transported across the outer chloroplast membrane by the TOC complex, or translocon on the outer chloroplast membrane.[59]
The chloroplast membranes sometimes protrude out into the cytoplasm, forming a stromule, or stroma-containing tubule. Stromules are very rare in chloroplasts, and are much more common in other plastids like chromoplasts and amyloplasts in petals and roots, respectively.[60][61] They may exist to increase the chloroplast's surface area for cross-membrane transport, connect two or more chloroplasts allowing them to exchange metabolites, or both.[18]
Intermembrane space and peptidoglycan wall
[edit]
Usually, a thin intermembrane space about 10–20 nanometers thick exists between the outer and inner chloroplast membranes.[62]
Glaucophyte algal chloroplasts have a peptidoglycan layer between the chloroplast membranes. It corresponds to the peptidoglycan cell wall of their cyanobacterial ancestors, which is located between their two cell membranes. These chloroplasts are called muroplasts (from Latin "mura", meaning "wall"). Other chloroplasts have lost the cyanobacterial wall, leaving an intermembrane space between the two chloroplast envelope membranes.[18]
Inner chloroplast membrane
[edit]The inner chloroplast membrane borders the stroma and regulates passage of materials in and out of the chloroplast. After passing through the TOC complex in the outer chloroplast membrane, polypeptides must pass through the TIC complex (translocon on the inner chloroplast membrane) which is located in the inner chloroplast membrane.[59]
In addition to regulating the passage of materials, the inner chloroplast membrane is where fatty acids, lipids, and carotenoids are synthesized.[18]
Peripheral reticulum
[edit]Some chloroplasts contain a structure called the chloroplast peripheral reticulum.[62] It is often found in the chloroplasts of C4 plants, though it's also been found in some C3 angiosperms,[18] and even some gymnosperms.[63] The chloroplast peripheral reticulum consists of a maze of membranous tubes and vesicles continuous with the inner chloroplast membrane that extends into the internal stromal fluid of the chloroplast. Its purpose is thought to be to increase the chloroplast's surface area for cross-membrane transport between its stroma and the cell cytoplasm. The small vesicles sometimes observed may serve as transport vesicles to shuttle stuff between the thylakoids and intermembrane space.[64]
Stroma
[edit]The protein-rich,[18] alkaline,[57] aqueous fluid within the inner chloroplast membrane and outside of the thylakoid space is called the stroma,[18] which corresponds to the cytosol of the original cyanobacterium. Nucleoids of chloroplast DNA, chloroplast ribosomes, the thylakoid system with plastoglobuli, starch granules, and many proteins can be found floating around in it. The Calvin cycle, which fixes CO2 into sugar takes place in the stroma.
Chloroplast ribosomes
[edit]Chloroplasts have their own ribosomes, which they use to synthesize a small fraction of their proteins. Chloroplast ribosomes are about two-thirds the size of cytoplasmic ribosomes (around 17 nm vs 25 nm).[62] They take mRNAs transcribed from the chloroplast DNA and translate them into protein. While similar to bacterial ribosomes,[3] chloroplast translation is more complex than in bacteria, so chloroplast ribosomes include some chloroplast-unique features.[65]
Plastoglobuli
[edit]Plastoglobuli (singular plastoglobulus, sometimes spelled plastoglobule(s)), are spherical globules of lipids and proteins[18] about 10–15 nanometers across.[62] They are surrounded by a lipid monolayer.[66] Plastoglobuli are found in all chloroplasts,[62] but become more common when the chloroplast is under oxidative stress,[66] or when it ages and transitions into a gerontoplast.[18] They are also common in etioplasts, but decrease in number as the etioplasts mature into chloroplasts.[66]
Plastoglobuli were once thought to be free-floating in the stroma, but it is now thought that they are permanently attached either to a thylakoid or to another plastoglobulus attached to a thylakoid. Plastoglubuli contain both structural proteins and enzymes involved in lipid synthesis and metabolism. They can form when a bubble appears between the layers of the lipid bilayer of the thylakoid membrane, or bud from existing plastoglubuli—though they never detach and float off into the stroma.[66]
Starch granules
[edit]Starch granules are very common in chloroplasts, typically taking up 15% of the organelle's volume,[67] though in some other plastids like amyloplasts, they can be big enough to distort the shape of the organelle.[62] Starch granules are simply accumulations of starch in the stroma, and are not bounded by a membrane.[62]
Starch granules appear and grow throughout the day, as the chloroplast synthesizes sugars, and are consumed at night to fuel respiration and continue sugar export into the phloem,[68] though in mature chloroplasts, it's rare for a starch granule to be completely consumed or for a new granule to accumulate.[67]
Starch granules vary in composition and location across different chloroplast lineages. In red algae, starch granules are found in the cytoplasm rather than in the chloroplast.[69] In C4 plants, mesophyll chloroplasts, which do not synthesize sugars, lack starch granules.[18]
Rubisco
[edit]
The chloroplast stroma contains many proteins, though the most common and important is Rubisco, which is probably also the most abundant protein on the planet.[57] Rubisco is the enzyme that fixes CO2 into sugar molecules. In C3 plants, rubisco is abundant in all chloroplasts, though in C4 plants, it's confined to the bundle sheath chloroplasts, where the Calvin cycle is carried out in C4 plants.[70]
Pyrenoids
[edit]The chloroplasts of some hornworts[71] and algae contain structures called pyrenoids. They are not found in higher plants.[72] Pyrenoids are roughly spherical and highly refractive bodies which are a site of starch accumulation in plants that contain them. They consist of an matrix opaque to electrons, surrounded by two hemispherical starch plates. The starch is accumulated as the pyrenoids mature.[73] In algae with carbon concentrating mechanisms, the enzyme rubisco is found in the pyrenoids. Starch can also accumulate around the pyrenoids when CO2 is scarce.[72] Pyrenoids can divide to form new pyrenoids, or be produced "de novo".[73][74]
Thylakoid system
[edit]
Suspended within the chloroplast stroma is the thylakoid system, a highly dynamic collection of membranous sacks called thylakoids where chlorophyll is found and the light reactions of photosynthesis happen.[75] In most vascular plant chloroplasts, the thylakoids are arranged in stacks called grana,[76] though in certain C4 plant chloroplasts[70] and some algal chloroplasts, the thylakoids are free floating.[8]
Granal structure
[edit]Using a light microscope, it's just barely possible to see tiny green granules—which were named grana.[62] With electron microscopy, it became possible to see the thylakoid system in more detail, revealing it to consist of stacks of flat thylakoids which made up the grana, and long interconnecting stromal thylakoids which linked different grana.[62] In the transmission electron microscope, thylakoid membranes appear as alternating light-and-dark bands, 8.5 nanometers thick.[62]
For a long time, the three dimensional structure of the thylakoid system has been unknown or disputed. One model has the granum as a stack of thylakoids linked by helical stromal thylakoids; the other has the granum as a single folded thylakoid connected in a "hub and spoke" way to other grana by stromal thylakoids. While the thylakoid system is still commonly depicted according to the folded thylakoid model,[75] it was determined in 2011 that the stacked and helical thylakoids model is correct.[77]
In the helical thylakoid model, grana consist of a stack of flattened circular granal thylakoids that resemble pancakes. Each granum can contain anywhere from two to a hundred thylakoids,[62] though grana with 10–20 thylakoids are most common.[76] Wrapped around the grana are helicoid stromal thylakoids, also known as frets or lamellar thylakoids. The helices ascend at an angle of 20–25°, connecting to each granal thylakoid at a bridge-like slit junction. The helicoids may extend as large sheets that link multiple grana, or narrow to tube-like bridges between grana.[77] While different parts of the thylakoid system contain different membrane proteins, the thylakoid membranes are continuous and the thylakoid space they enclose form a single continuous labyrinth.[76]
Thylakoids
[edit]Thylakoids (sometimes spelled thylakoïds),[78] are small interconnected sacks which contain the membranes that the light reactions of photosynthesis take place on. The word thylakoid comes from the Greek word thylakos which means "sack".[79]
Embedded in the thylakoid membranes are important protein complexes which carry out the light reactions of photosynthesis. Photosystem II and photosystem I contain light-harvesting complexes with chlorophyll and carotenoids that absorb light energy and use it to energize electrons. Molecules in the thylakoid membrane use the energized electrons to pump hydrogen ions into the thylakoid space, decreasing the pH and turning it acidic. ATP synthase is a large protein complex that harnesses the concentration gradient of the hydrogen ions in the thylakoid space to generate ATP energy as the hydrogen ions flow back out into the stroma—much like a dam turbine.[57]
There are two types of thylakoids—granal thylakoids, which are arranged in grana, and stromal thylakoids, which are in contact with the stroma. Granal thylakoids are pancake-shaped circular disks about 300–600 nanometers in diameter. Stromal thylakoids are helicoid sheets that spiral around grana.[76] The flat tops and bottoms of granal thylakoids contain only the relatively flat photosystem II protein complex. This allows them to stack tightly, forming grana with many layers of tightly appressed membrane, called granal membrane, increasing stability and surface area for light capture.[76]
In contrast, photosystem I and ATP synthase are large protein complexes which jut out into the stroma. They can't fit in the appressed granal membranes, and so are found in the stromal thylakoid membrane—the edges of the granal thylakoid disks and the stromal thylakoids. These large protein complexes may act as spacers between the sheets of stromal thylakoids.[76]
The number of thylakoids and the total thylakoid area of a chloroplast is influenced by light exposure. Shaded chloroplasts contain larger and more grana with more thylakoid membrane area than chloroplasts exposed to bright light, which have smaller and fewer grana and less thylakoid area. Thylakoid extent can change within minutes of light exposure or removal.[64]
Pigments and chloroplast colors
[edit]Inside the photosystems embedded in chloroplast thylakoid membranes are various photosynthetic pigments, which absorb and transfer light energy. The types of pigments found are different in various groups of chloroplasts, and are responsible for a wide variety of chloroplast colorations.
Chlorophylls
[edit]Chlorophyll a is found in all chloroplasts, as well as their cyanobacterial ancestors. Chlorophyll a is a blue-green pigment[80] partially responsible for giving most cyanobacteria and chloroplasts their color. Other forms of chlorophyll exist, such as the accessory pigments chlorophyll b, chlorophyll c, chlorophyll d,[8] and chlorophyll f.
Chlorophyll b is an olive green pigment found only in the chloroplasts of plants, green algae, any secondary chloroplasts obtained through the secondary endosymbiosis of a green alga, and a few cyanobacteria.[8] It's the chlorophylls a and b together that make most plant and green algal chloroplasts green.[80]
Chlorophyll c is mainly found in secondary endosymbiotic chloroplasts that originated from a red alga, though it's not found in chloroplasts of red algae themselves. Chlorophyll c is also found in some green algae and cyanobacteria.[8]
Chlorophylls d and f are pigments found only in some cyanobacteria.[8][81]
Carotenoids
[edit]![Delesseria sanguinea, a red alga, has chloroplasts that contain red pigments like phycoerytherin that mask their blue-green chlorophyll a.[20]](http://upload.wikimedia.org/wikipedia/commons/thumb/1/19/Delesseria_sanguinea_Helgoland.JPG/300px-Delesseria_sanguinea_Helgoland.JPG)
In addition to chlorophylls, another group of yellow–orange[80] pigments called carotenoids are also found in the photosystems. There are about thirty photosynthetic carotenoids.[82] They help transfer and dissipate excess energy,[8] and their bright colors sometimes override the chlorophyll green, like during the fall, when the leaves of some land plants change color.[83] β-carotene is a bright red-orange carotenoid found in nearly all chloroplasts, like chlorophyll a.[8] Xanthophylls, especially the orange-red zeaxanthin, are also common.[82] Many other forms of carotenoids exist that are only found in certain groups of chloroplasts.[8]
Phycobillins
[edit]Phycobillins are a third group of pigments found in cyanobacteria, and glaucophyte and red algal chloroplasts.[8] Phycobillins come in all colors, though phycoerytherin is one of the pigments that makes many red algae red.[84] Phycobillins often organize into relatively large protein complexes about 40 nanometers across called phycobilisomes.[8] Like photosystem I and ATP synthase, phycobillisomes jut into the stroma, preventing thylakoid stacking in red algal chloroplasts.[8] Cryptophyte chloroplasts and some cyanobacteria don't have their phycobilin pigments organized into phycobillisomes, and keep them in their thylakoid space instead.[8]
Specialized chloroplasts in C4 plants
[edit]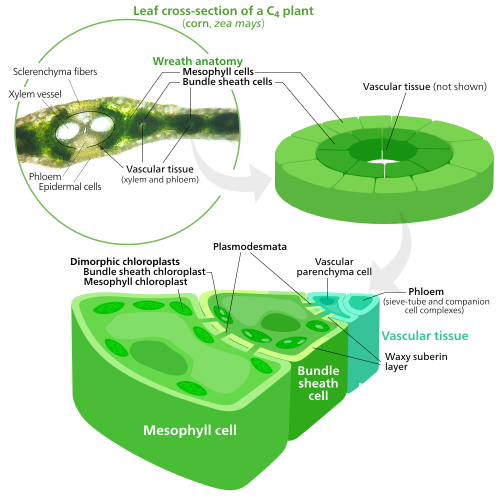
To fix carbon dioxide into sugar molecules in the process of photosynthesis, chloroplasts use an enzyme called rubisco. Rubisco has a problem—it has trouble distinguishing between carbon dioxide and oxygen, so at high oxygen concentrations, rubisco starts accidentally adding oxygen to sugar precursors. This has the end result of ATP energy being wasted and CO2 being released, all with no sugar being produced. This is a big problem, since O2 is produced by the initial light reactions of photosynthesis, causing issues down the line in the Calvin cycle which uses rubisco.[85]
C4 plants evolved a way to solve this—by spatially separating the light reactions and the Calvin cycle. The light reactions, which store light energy in ATP and NADPH, are done in the mesophyll cells of a C4 leaf. The Calvin cycle, which uses the stored energy to make sugar using rubisco, is done in the bundle sheath cells, a layer of cells surrounding a vein in a leaf.[85]
As a result, chloroplasts in C4 mesophyll cells and bundle sheath cells are specialized for each stage of photosynthesis. In mesophyll cells, chloroplasts are specialized for the light reactions, so they lack rubisco, and have normal grana and thylakoids,[70] which they use to make ATP and NADPH, as well as oxygen. They store CO2 in a four-carbon compound, which is why the process is called C4 photosynthesis. The four-carbon compound is then transported to the bundle sheath chloroplasts, where it drops off CO2 and returns to the mesophyll. Bundle sheath chloroplasts do not carry out the light reactions, preventing oxygen from building up in them and disrupting rubisco activity.[85] Because of this, they lack thylakoids organized into grana stacks—though bundle sheath chloroplasts still have free-floating thylakoids in the stroma where they still carry out cyclic electron flow, a light-driven method of synthesizing ATP to power the Calvin cycle without generating oxygen. They lack photosystem II, and only have photosystem I—the only protein complex needed for cyclic electron flow.[70][85] Because the job of bundle sheath chloroplasts is to carry out the Calvin cycle and make sugar, they often contain large starch grains.[70]
Both types of chloroplast contain large amounts of chloroplast peripheral reticulum,[70] which they use to get more surface area to transport stuff in and out of them.[63][64] Mesophyll chloroplasts have a little more peripheral reticulum than bundle sheath chloroplasts.[86]
Distribution in a plant
[edit]Not all cells in a multicellular plant contain chloroplasts. All green parts of a plant contain chloroplasts—the chloroplasts, or more specifically, the chlorophyll in them are what make the photosynthetic parts of a plant green.[75] The plant cells which contain chloroplasts are usually parenchyma cells, though chloroplasts can also be found in collenchyma tissue.[87] A plant cell which contains chloroplasts is known as a chlorenchyma cell. A typical chlorenchyma cell of a land plant contains about 10 to 100 chloroplasts.

In some plants such as cacti, chloroplasts are found in the stems,[88] though in most plants, chloroplasts are concentrated in the leaves. One square millimeter of leaf tissue can contain half a million chloroplasts.[75] Within a leaf, chloroplasts are mainly found in the mesophyll layers of a leaf, and the guard cells of stomata. Palisade mesophyll cells can contain 30–70 chloroplasts per cell, while stomatal guard cells contain only around 8–15 per cell, as well as much less chlorophyll. Chloroplasts can also be found in the bundle sheath cells of a leaf, especially in C4 plants, which carry out the Calvin cycle in their bundle sheath cells. They are often absent from the epidermis of a leaf.[89]
Algal chloroplasts
[edit]This section needs expansion. You can help by adding to it. (May 2013) |
In the cells of many alga there is only one chloroplast[90] (for example in Chlorella, it fills much of the cell and is bell-shaped).
Cellular location
[edit]Chloroplast movement
[edit]The chloroplasts of plant and algal cells can orient themselves to best suit the available light. In low-light conditions, they will spread out in a sheet—maximizing the surface area to absorb light. Under intense light, they will seek shelter by aligning in vertical columns along the plant cell's cell wall or turning sideways so that light strikes them edge-on. This reduces exposure and protects them from photooxidative damage.[91] This ability to distribute chloroplasts so that they can take shelter behind each other or spread out may be the reason why land plants evolved to have many small chloroplasts instead of a few big ones.[92] Chloroplast movement is considered one of the most closely regulated stimulus-response systems that can be found in plants.[93] Mitochondria have also been observed to follow chloroplasts as they move.[94]
In higher plants, chloroplast movement is run by phototropins, blue light photoreceptors also responsible for plant phototropism. In some algae, mosses, ferns, and flowering plants, chloroplast movement is influenced by red light in addition to blue light,[91] though very long red wavelengths inhibit movement rather than speeding it up. Blue light generally causes chloroplasts to seek shelter, while red light draws them out to maximize light absorption.[94]
Studies of Vallisneria gigantea, an aquatic flowering plant, have shown that chloroplasts can get moving within five minutes of light exposure, though they don't initially show any net directionality. They may move along microfilament tracks, and the fact that the microfilament mesh changes shape to form a honeycomb structure surrounding the chloroplasts after they have moved suggests that microfilaments may help to anchor chloroplasts in place.[93][94]
Role in plant immunity
[edit]Plants lack specialized immune cells—all plant cells participate in the plant immune response. Chloroplasts, along with the nucleus, cell membrane, and endoplasmic reticulum,[95] are key players in pathogen defense. Due to its role in a plant cell's immune response, pathogens frequently target the chloroplast.[95]
Plants have two main immune responses—the hypersensitive response, in which infected cells seal themselves off and undergo programmed cell death, and systemic acquired resistance, where infected cells release signals warning the rest of the plant of a pathogen's presence. Chloroplasts stimulate both responses by purposely damaging their photosynthetic system, producing reactive oxygen species. High levels of reactive oxygen species will cause the hypersensitive response. The reactive oxygen species also directly kill any pathogens within the cell. Lower levels of reactive oxygen species initiate systemic acquired resistance, triggering defense-molecule production in the rest of the plant.[95]
In some plants, chloroplasts are known to move closer to the infection site and the nucleus during an infection.[95]
Chloroplasts can serve as cellular sensors. After detecting stress in a cell, which might be due to a pathogen, chloroplasts begin producing molecules like salicylic acid, jasmonic acid, nitric oxide and reactive oxygen species which can serve as defense-signals. As cellular signals, reactive oxygen species are unstable molecules, so they probably don't leave the chloroplast, but instead pass on their signal to an unknown second messenger molecule. All these molecules initiate retrograde signaling—signals from the chloroplast that regulate gene expression in the nucleus.[95]
In addition to defense signaling, chloroplasts, with the help of the peroxisomes,[96] help synthesize an important defense molecule, jasmonate. Chloroplasts synthesize all the fatty acids in a plant cell[95][97]—linoleic acid, a fatty acid, is a precursor to jasmonate.[95]
Photosynthetic chemistry
[edit]One of the most important function of the chloroplast is carrying out photosynthesis to make food in the form of sugars for an alga or plant. Water (H2O) and carbon dioxide (CO2) are used in photosynthesis, and sugar and oxygen (O2) is made, using light energy. Photosynthesis is divided into two stages—the light reactions, where water is split to produce oxygen, and the dark reactions, or Calvin cycle, which builds sugar molecules from carbon dioxide. The two phases are linked by the energy carriers adenosine triphosphate (ATP) and nicotinamide adenine dinucleotide phosphate (NADP+).[98]
Light reactions
[edit]
The light reactions take place on the thylakoid membranes. They take light energy and store it in NADPH, a form of NADP+, and ATP to fuel the dark reactions.
Energy carriers
[edit]ATP is the phosphorylated version of adenosine diphosphate (ADP), which stores energy in a cell and powers most cellular activities. ATP is the energized form, while ADP is the (partially) depleted form. NADP+ is an electron carrier which ferries high energy electrons. In the light reactions, it gets reduced, meaning it picks up electrons, becoming NADPH.
Photophosphorylation
[edit]Like mitochondria, chloroplasts use the potential energy stored in an H+, or hydrogen ion gradient to generate ATP energy. The two photosystems capture light energy to energize electrons taken from water, and release them down an electron transport chain. The molecules between the photosystems harness the electrons' energy to pump hydrogen ions into the thylakoid space, creating a concentration gradient, with more hydrogen ions (up to a thousand times as many)[57] inside the thylakoid system than in the stroma. The hydrogen ions in the thylakoid space then diffuse back down their concentration gradient, flowing back out into the stroma through ATP synthase. ATP synthase uses the energy from the flowing hydrogen ions to phosphorylate adenosine diphosphate into adenosine triphosphate, or ATP.[57] Because chloroplast ATP synthase projects out into the stroma, the ATP is synthesized there, in position to be used in the dark reactions.[99]
NADP+ reduction
[edit]Electrons are often removed from the electron transport chains to charge NADP+ with electrons, reducing it to NADPH. Like ATP synthase, ferredoxin-NADP+ reductase, the enzyme that reduces NADP+, releases the NADPH it makes into the stroma, right where it's needed for the dark reactions.[99]
Because NADP+ reduction removes electrons from the electron transport chains, they must be replaced—the job of photosystem II, which splits water molecules (H2O) to obtain the electrons from its hydrogen atoms.[57][98]
Cyclic photophosphorylation
[edit]While photosystem II photolyzes water to obtain and energize new electrons, photosystem I simply reenergizes depleted electrons at the end of an electron transport chain. Normally, the reenergized electrons are taken by NADP+, though sometimes they can flow back down more H+-pumping electron transport chains to transport more hydrogen ions into the thylakoid space to generate more ATP. This is termed cyclic photophosphorylation because the electrons are recycled. Cyclic photophosphorylation is common in C4 plants, which need more ATP than NADPH.[85]
Dark reactions
[edit]The Calvin cycle, also known as the dark reactions, is a series of biochemical reactions that fixes CO2 into G3P sugar molecules and uses the energy and electrons from the ATP and NADPH made in the light reactions. The Calvin cycle takes place in the stroma of the chloroplast.[85]
While named "the dark reactions", in most plants, they take place in the light, since the dark reactions are dependent on the products of the light reactions.[75]
Carbon fixation and G3P synthesis
[edit]The Calvin cycle starts by using the enzyme Rubisco to fix CO2 into five-carbon Ribulose bisphosphate (RuBP) molecules. The result is unstable six-carbon molecules that immediately break down into three-carbon molecules called 3-phosphoglyceric acid, or 3-PGA. The ATP and NADPH made in the light reactions is used to convert the 3-PGA into glyceraldehyde-3-phosphate, or G3P sugar molecules. Most of the G3P molecules are recycled back into RuBP using energy from more ATP, but one out of every six produced leaves the cycle—the end product of the dark reactions.[85]
Sugar synthesis
[edit]Glyceraldehyde-3-phosphate can double up and form glucose-1-phosphate, glucose-6-phosphate or fructose-6-phosphate molecules which each include a phosphate group. To synthesize sucrose, a disaccharide commonly known as table sugar, the G3P molecules are first transported into the cytoplasm by a translocon in the chloroplast membrane. In the cytoplasm, they double up to form fructose-6-phosphate, join with glucose monomers, and have their phosphate groups removed to become the disaccharide sucrose.[100]
Alternatively, glucose monomers in the chloroplast can be linked together to make starch, which accumulates into starch grains in the chloroplast.[100]
Under conditions such as high atmospheric CO2 concentrations, large starch grains may form in the chloroplasts, distorting the grana and thylakoids. The starch granules displace the thylakoids, but leave them intact.[101]
Waterlogged roots can also cause starch buildup in the chloroplasts, possibly due to less sugar being exported through the phloem. This depletes a plant's free phosphate supply, which indirectly stimulates chloroplast starch synthesis.[101] While linked to low photosynthesis rates, the starch grains themselves may not necessarily interfere significantly with the efficiency of photosynthesis,[102] and might simply be a side effect of another photosynthesis-depressing factor.[101]
Photorespiration
[edit]Photorespiration can occur when the oxygen concentration is too high. Rubisco cannot distinguish between oxygen and carbon dioxide very well, so it can accidentally add O2 instead of CO2 to RuBP. This process reduces the efficiency of photosynthesis—it consumes ATP and oxygen, releases CO2, and produces no sugar. It can waste up to half the carbon fixed by the Calvin cycle.[98] Several mechanisms have evolved in different lineages that raise the carbon dioxide concentration relative to oxygen within the chloroplast, increasing the efficiency of photosynthesis. These mechanisms are called carbon dioxide concentrating mechanisms, or CCMs. These include Crassulacean acid metabolism, C4 carbon fixation,[98] and pyrenoids. Chloroplasts in C4 plants are notable as they exhibit a distinct chloroplast dimorphism.
pH
[edit]Because of the H+ gradient across the thylakoid membrane, the interior of the thylakoid is acidic, with a pH around 4,[103] while the stroma is slightly basic, with a pH of around 8.[104] The optimal stroma pH for the Calvin cycle is 8.1, with the reaction nearly stopping when the pH falls below 7.3.[105]
CO2 in water can form carbonic acid, which can disturb the pH of isolated chloroplasts, interfering with photosynthesis, even though CO2 is used in photosynthesis. However, chloroplasts in living plant cells are not affected by this as much.[104]
Chloroplasts can pump K+ and H+ ions in and out of themselves using a poorly understood light-driven transport system.[104]
In the presence of light, the pH of the thylakoid lumen can drop up to 1.5 pH units, while the pH of the stroma can rise by nearly one pH unit.[105]
Other chemistry
[edit]Other chemical products
[edit]This section needs expansion with: needs more about lipids, also paramylon. You can help by adding to it. (March 2013) |
Chloroplasts are the site of complex lipid metabolism.[106]
Development and differentiation
[edit]Chloroplasts are a special type of plant cell organelle called a plastid, though the two terms are sometimes used interchangeably. There are many other types of plastids, which carry out various functions. All chloroplasts in a plant are descended from undifferentiated proplastids found in the zygote,[107] or fertilized egg. Proplastids are commonly found in an adult plant's apical meristems. Chloroplasts do not normally develop from proplastids in root tip meristems[108]—instead, the formation of starch-storing amyloplasts is more common.[107]
In shoots, proplastids from shoot apical meristems gradually develop into chloroplasts in photosynthetic leaf tissues as the leaf matures, if exposed to the required light. This process involves invaginations of the inner plastid membrane, forming sheets of membrane that project into the internal stroma. These membrane sheets then fold to form thylakoids and grana.[109]
If angiosperm shoots are not exposed to the required light for chloroplast formation, proplastids may develop into an etioplast stage before becoming chloroplasts. An etioplast is a plastid that lacks chlorophyll, and has inner membrane invaginations that form a lattice of tubes in their stroma, called a prolamellar body. Within a few minutes of light exposure, the prolamellar body begins to reorganize into stacks of thylakoids, and chlorophyll starts to be produced. This process, where the etioplast becomes a chloroplast, takes several hours.[109] Gymnosperms do not require light to form chloroplasts.[109]
Light, however, does not guarantee that a proplastid will develop into a chloroplast—what type of plastid a proplastid becomes is largely influenced by the kind of cell it resides in.[107]
Plastid interconversion
[edit]Plastid differentiation is not permanent, in fact many interconversions are possible. Chloroplasts may be converted to chromoplasts, which are pigment-filled plastids responsible for the bright colors you see in flowers and ripe fruit. Starch storing amyloplasts can also be converted to chromoplasts, and it's possible for proplastids to develop straight into chromoplasts. Chromoplasts and amyloplasts can also become chloroplasts, like what happens when you illuminate a carrot or a potato. If a plant is injured, or something else causes a plant cell to revert to a meristematic state, chloroplasts and other plastids can turn back into proplastids. Chloroplast, amyloplast, chromoplast, proplast, etc, are not absolute states—intermediate forms are common.[107]
Chloroplast division
[edit]This section needs expansion with: functions, Z-ring dynamic assembly, regulators such as Giant Chloroplast 1. You can help by adding to it. (February 2013) |
Most chloroplasts in a photosynthetic cell do not develop directly from proplastids or etioplasts. In fact, a typical shoot meristematic plant cell contains only 7–20 proplastids. These proplastids differentiate into chloroplasts, which divide to create the 30–70 chloroplasts found in a mature photosynthetic plant cell. If the cell divides, chloroplast division provides the additional chloroplasts to partition between the two daughter cells.[110]
In single-celled algae, chloroplast division is the only way new chloroplasts are formed. There is no proplastid differentiation—when an algal cell divides, its chloroplast divides along with it, and each daughter cell receives a mature chloroplast.[109]
Almost all chloroplasts in a cell divide, rather than a small group of rapidly dividing chloroplasts.[111] Chloroplasts have no definite S-phase—their DNA replication is not synchronized or limited to that of their host cells.[112] Much of what we know about chloroplast division comes from studying the alga Cyanidioschyzon merolæ.[92]
The division process starts when the proteins FtsZ1 and FtsZ2 assemble into filaments, and with the help of a protein ARC6, form a structure called a Z-ring within the chloroplast's stroma.[92][113] The Min system manages the placement of the Z-ring, ensuring that the chloroplast is cleaved more or less evenly. The protein MinD prevents FtsZ from linking up and forming filaments. Another protein ARC3 may also be involved, but it is not very well understood. These proteins are active at the poles of the chloroplast, preventing Z-ring formation there, but near the center of the chloroplast, MinE inhibits them, allowing the Z-ring to form.[92]
Next, the two plastid-dividing rings, or PD rings form. The inner plastid-dividing ring is located in the inner side of the chloroplast's inner membrane, and is formed first.[92] The outer plastid-dividing ring is found wrapped around the outer chloroplast membrane. It consists of filaments about 5 nanometers across,[92] arranged in rows 6.4 nanometers apart, and shrinks to squeeze the chloroplast. This is when chloroplast constriction begins.[113]
In a few species like Cyanidioschyzon merolæ, chloroplasts have a third plastid-dividing ring located in the chloroplast's intermembrane space.[92][113]
Late into the constriction phase, dynamin proteins assemble around the outer plastid-dividing ring,[113] helping provide force to squeeze the chloroplast.[92] Meanwhile, the Z-ring and the inner plastid-dividing ring break down.[113] During this stage, the many chloroplast DNA plasmids floating around in the stroma are partitioned and distributed to the two forming daughter chloroplasts.[114]
Later, the dynamins migrate under the outer plastid dividing ring, into direct contact with the chloroplast's outer membrane,[113] to cleave the chloroplast in two daughter chloroplasts.[92]
A remnant of the outer plastid dividing ring remains floating between the two daughter chloroplasts, and a remnant of the dynamin ring remains attached to one of the daughter chloroplasts.[113]
Of the five or six rings involved in chloroplast division, only the outer plastid-dividing ring is present for the entire constriction and division phase—while the Z-ring forms first, constriction does not begin until the outer plastid-dividing ring forms.[113]
Regulation
[edit]In species of algae which contain a single chloroplast, regulation of chloroplast division is extremely important to ensure that each daughter cell receives a chloroplast. In organisms like plants, whose cells contain multiple chloroplasts, coordination is looser and less important. It's likely that chloroplast and cell division are somewhat synchronized, though the mechanisms for it are mostly unknown.[92]
Light has been shown to be a requirement for chloroplast division. Chloroplasts can grow and progress through some of the constriction stages under poor quality green light, but are slow to complete division—they require exposure to bright white light to complete division. Spinach leaves grown under green light have been observed to contain many large dumbbell-shaped chloroplasts. Exposure to white light can stimulate these chloroplasts to divide and reduce the population of dumbbell-shaped chloroplasts.[111][114]
Chloroplast inheritance
[edit]Like mitochondria, chloroplasts are usually inherited from a single parent. Biparental chloroplast inheritance—where plastid genes are inherited from both parent plants—occurs in very low levels in some flowering plants.[115]
Many mechanisms prevent biparental chloroplast DNA inheritance including selective destruction of chloroplasts or their genes within the gamete or zygote, and chloroplasts from one parent being excluded from the embryo. Parental chloroplasts can be sorted so that only one type is present in each offspring.[116]
Gymnosperms, such as pine trees, mostly pass on chloroplasts paternally,[117] while flowering plants often inherit chloroplasts maternally.[118][119] Flowering plants were once thought to only inherit chloroplasts maternally. However, there are now many documented cases of angiosperms inheriting chloroplasts paternally.[115]
Angiosperms which pass on chloroplasts maternally have many ways to prevent paternal inheritance. Most of them produce sperm cells which do not contain any plastids. There are many other documented mechanisms that prevent paternal inheritance in these flowering plants, such as different rates of chloroplast replication within the embryo.[115]
Among angiosperms, paternal chloroplast inheritance is observed more often in hybrids than in offspring from parents of the same species. This suggests that incompatible hybrid genes might interfere with the mechanisms that prevent paternal inheritance.[115]
Transplastomic plants
[edit]Recently, chloroplasts have caught attention by developers of genetically modified crops. Since in most flowering plants, chloroplasts are not inherited from the male parent, transgenes in these plastids cannot be disseminated by pollen. This makes plastid transformation a valuable tool for the creation and cultivation of genetically modified plants that are biologically contained, thus posing significantly lower environmental risks. This biological containment strategy is therefore suitable for establishing the coexistence of conventional and organic agriculture. While the reliability of this mechanism has not yet been studied for all relevant crop species, recent results in tobacco plants are promising, showing a failed containment rate of transplastomic plants at 3 in 1,000,000.[119]
See also
[edit]Notes
[edit] This article incorporates public domain material from Science Primer. NCBI. Archived from the original on 8 December 2009.
This article incorporates public domain material from Science Primer. NCBI. Archived from the original on 8 December 2009.
References
[edit]- ^ Campbell, Neil A. (2006). Biology: Exploring Life. Boston, Massachusetts: Pearson Prentice Hall. ISBN 978-0-13-250882-7.
{{cite book}}: Unknown parameter|coauthors=ignored (|author=suggested) (help) - ^ "chloroplast". Online Etymology Dictionary.
- ^ a b c d e f Biology 8th Edition Campbell & Reece. Benjamin Cummings (Pearson). 2009. p. 516.
- ^ Mereschkowsky C (1905). "Über Natur und Ursprung der Chromatophoren im Pflanzenreiche". Biol Centralbl. 25: 593–604.
- ^ Schimper AFW (1883). "Über die Entwicklung der Chlorophyllkörner und Farbkörper". Bot. Zeitung. 41: 105–14, 121–31, 137–46, 153–62.
- ^ a b Kumar, K.; Mella-Herrera, R. A.; Golden, J. W. (April 2010). "Cyanobacterial Heterocysts". Cold Spring Harb Perspect Biol. 2 (4): a000315. doi:10.1101/cshperspect.a000315. PMC 2845205. PMID 20452939.
{{cite journal}}: CS1 maint: date and year (link) - ^ a b c d McFadden, Geoffrey I.; Van Dooren, Giel G. (NaN undefined NaN). "Evolution: Red Algal Genome Affirms a Common Origin of All Plastids". Current Biology. 14 (13): R514–R516. doi:10.1016/j.cub.2004.06.041. PMID 15242632. S2CID 18131616. Retrieved 7 April 2013.
{{cite journal}}: Check date values in:|date=(help) - ^ a b c d e f g h i j k l m n o p q r s t u v w x y z aa ab ac ad ae af ag ah ai aj ak al am Aronsson, edited by David G. Robinson, Anna Stina Sandelius, Henrik (2009). The Chloroplast Interactions with the Environment (PDF) ([Online-Ausg.] ed.). Berlin, Heidelberg: Springer Berlin Heidelberg. pp. 3–7. ISBN 9783540686965.
{{cite book}}:|first=has generic name (help)CS1 maint: multiple names: authors list (link) - ^ "Chloroplasts". Kimball's Biology Pages. Retrieved 27 December 2012.
- ^ Joyard J Block MA, Douce R (1991). "Molecular aspects of plastid envelope biochemistry". Eur J Biochem. 199 (3): 489–509. doi:10.1111/j.1432-1033.1991.tb16148.x. PMID 1868841.
- ^ "Chloroplast". Encyclopedia of Science. Retrieved 27 December 2012.
- ^ a b c d e f g h i j k l m n o p q r s t Keeling, Patrick J (October 2004). "Diversity and evolutionary history of plastids and their hosts". American Journal of Botany. 91 (10): 1481–93. doi:10.3732/ajb.91.10.1481. PMID 21652304. S2CID 17522125. Retrieved 27 January 2013.
{{cite journal}}: CS1 maint: date and year (link) - ^ a b c d e f Nakayama, Takuro; Archibald, John M. (April 2012). "Evolving a photosynthetic organelle". BMC Biology. 10: 35. doi:10.1186/1741-7007-10-35. PMC 3337241. PMID 22531210.
{{cite journal}}: CS1 maint: date and year (link) CS1 maint: unflagged free DOI (link) - ^ a b c d e f McFadden, Geoffrey I (January 2001). "Chloroplast Origin and Integration". Plant Physiology. 125 (1): 50–3. doi:10.1104/pp.125.1.50. PMC 1539323. PMID 11154294. Retrieved 27 January 2013.
{{cite journal}}: CS1 maint: date and year (link) - ^ Archibald, J.M. (2009). "The Puzzle of Plastid Evolution". Current Biology. 19 (2): R81–R88. doi:10.1016/j.cub.2008.11.067. PMID 19174147. S2CID 51989.
- ^ a b Ball, Steven; Colleoni, Christophe; Cenci, Ugo; Raj, Jenifer Nirmal; Tirtiaux, Catherine (10 January 2011). "The evolution of glycogen and starch metabolism in eukaryotes gives molecular clues to understand the establishment of plastid endosymbiosis". Journal of Experimental Botany. 62 (6): 1775–1801. doi:10.1093/jxb/erq411. PMID 21220783. Retrieved 6 June 2013.
- ^ Nair, Sethu C.; Striepen, Boris (30). "What Do Human Parasites Do with a Chloroplast Anyway?". PLOS Biology. 9 (8): e1001137. doi:10.1371/journal.pbio.1001137. PMC 3166169. PMID 21912515.
{{cite journal}}: Check date values in:|date=and|year=/|date=mismatch (help); Unknown parameter|month=ignored (help)CS1 maint: unflagged free DOI (link) - ^ a b c d e f g h i j k l m n o p q r s t u v w Wise, edited by Robert R. (2006). The structure and function of plastids. Dordrecht: Springer. pp. 3–21. ISBN 978-1-4020-4061-0.
{{cite book}}:|first=has generic name (help); Unknown parameter|coauthors=ignored (|author=suggested) (help) - ^ a b c d e f g h i j k l m Keeling, P. J. (2 February 2010). "The endosymbiotic origin, diversification and fate of plastids". Philosophical Transactions of the Royal Society B: Biological Sciences. 365 (1541): 729–748. doi:10.1098/rstb.2009.0103. PMC 2817223. PMID 20124341. Retrieved 8 June 2013.
- ^ a b c d e f g h i Biology 8th Edition Campbell & Reece. Benjamin Cummings (Pearson). 2009. pp. 582–592.
- ^ "rhodo-". The Free Dictionary. Farlex. Retrieved 7 June 2013.
- ^ a b Lewis, Louise A.; McCourt, Richard M. (1 October 2004). "Green algae and the origin of land plants". American Journal of Botany. 91 (10): 1535–1556. doi:10.3732/ajb.91.10.1535. PMID 21652308. Retrieved 7 June 2013.
- ^ Moroney, J. V.; Somanchi, A. (January 1999). "How Do Algae Concentrate CO2 to Increase the Efficiency of Photosynthetic Carbon Fixation?". Plant Physiology. 119 (1): 9–16. doi:10.1104/pp.119.1.9. PMC 1539202. PMID 9880340. Retrieved 7 June 2013.
{{cite journal}}: CS1 maint: date and year (link) - ^ a b Chaal, Balbir K.; Green, Beverley R. (February 2005). "Protein import pathways in 'complex' chloroplasts derived from secondary endosymbiosis involving a red algal ancestor". Plant Molecular Biology. 57 (3): 333–42. doi:10.1007/s11103-004-7848-y. PMID 15830125. S2CID 22619029.
{{cite journal}}: CS1 maint: date and year (link) - ^ Dorrell, Richard G.; Smith, Alison G. (July 2011). "Do Red and Green Make Brown?: Perspectives on Plastid Acquisitions within Chromalveolates". Eukaryot Cell. 10 (7): A SYMPHONY OF RED, GREEN, AND BROWN–THE DIVERSITY OF ALGAE. doi:10.1128/EC.00326-10. PMC 3147421. PMID 21622904.
{{cite journal}}: CS1 maint: date and year (link) - ^ Patrick J. Keeling (2004). "Diversity and evolutionary history of plastids and their hosts". American Journal of Botany. 91 (10): 1481–1493. doi:10.3732/ajb.91.10.1481. PMID 21652304. S2CID 17522125.[dead link]
- ^ a b c d Nowack, E. C. M.; Vogel, H.; Groth, M.; Grossman, A. R.; Melkonian, M.; Glockner, G. (January 2011). "Endosymbiotic gene transfer and transcriptional regulation of transferred genes in Paulinella chromatophora". Molecular Biology Evol. 28 (1): 407–22. doi:10.1093/molbev/msq209. PMID 20702568.
{{cite journal}}: CS1 maint: date and year (link) - ^ a b c d e Dann, Leighton (2002). Bioscience—Explained (PDF). Green DNA: BIOSCIENCE EXPLAINED.
- ^ a b c d e Clegg, M. T.; Gaut, B. S.; Learn, G. H.; Morton, B. R. (1994). "Rates and patterns of chloroplast DNA evolution". Proceedings of the National Academy of Sciences. 91 (15): 6795–6801. doi:10.1073/pnas.91.15.6795. PMC 44285. PMID 8041699.
- ^ a b c d e f Shaw, Joey; Lickey, Edgar B.; Schilling, Edward E.; Small, Randall L. (2007). "Comparison of whole chloroplast genome sequences to choose noncoding regions for phylogenetic studies in angiosperms: the tortoise and the hare III". American Journal of Botany. 94 (3): 275–88. doi:10.3732/ajb.94.3.275. PMID 21636401. S2CID 30501148. Retrieved 2 January 2013.
- ^ Skovgaard A (1998). "Role of chloroplast retention in a marine dinoflagellate" (PDF). Aquatic Microbial Ecology. 15: 293–301. doi:10.3354/ame015293. Archived from the original on 18 November 2010.
- ^ C.Michael Hogan. 2010. Deoxyribonucleic acid. Encyclopedia of Earth. National Council for Science and the Environment. eds. S.Draggan and C.Cleveland. Washington DC
- ^ "ctDNA — chloroplast DNA". AllAcronyms.com. Retrieved 2 January 2013.
- ^ The Oxford Dictionary of Abbreviations. ctDNA—Dictionary definition. 1998.
{{cite book}}: CS1 maint: location missing publisher (link) - ^ "Chloroplasts and Other Plastids". University of Hamburg. Retrieved 27 December 2012.
- ^ a b c Sandelius, Anna Stina (2009). The Chloroplast: Interactions with the Environment. Springer. p. 18. ISBN 9783540686965.
- ^ a b Burgess, Jeremy (1989). An introduction to plant cell development ([Pbk. ed.). Cambridge: Cambridge university press. p. 62. ISBN 0521316111.
- ^ a b c Kolodner, Richard; Tewari, K. K. (1979). "Inverted repeats in chloroplast DNA from higher plants". Proceedings of the National Academy of Sciences. 76 (1): 41–45. doi:10.1073/pnas.76.1.41. PMC 382872. PMID 16592612.
- ^ a b Palmer, Jeffrey D.; Thompson, William F. (1982). "Chloroplast DNA rearrangements are more frequent when a large inverted repeat sequence is lost". Cell. 29 (2): 537–50. doi:10.1016/0092-8674(82)90170-2. PMID 6288261. S2CID 11571695. Retrieved 2 January 2013.
- ^ Plant Biochemistry, 3rd edition. Academic Press. 2005. p. 517. ISBN 9780120883912.
- ^ a b Kobayashi, Tamaki; Takahara, Manabu; Miyagishima, Shin-ya; Kuroiwa, Haruko; Sasaki, Narie; Ohta, Niji; Matsuzaki, Motomichi; Kuroiwa, Tsuneyoshi (July 2002). "Detection and Localization of a Chloroplast-Encoded HU-Like Protein That Organizes Chloroplast Nucleoids". The Plant Cell. 14 (7): 1579–1589. doi:10.1105/tpc.002717. PMC 150708. PMID 12119376. Retrieved 21 April 2013.
{{cite journal}}: CS1 maint: date and year (link) - ^ a b c d Harris, E. H.; Boynton, J. E.; Gillham, N. W. (December 1994). "Chloroplast ribosomes and protein synthesis". American Society for Microbiology—Mol. Biol. Rev. 58 (4): 700–754. doi:10.1128/mr.58.4.700-754.1994. PMC 372988. PMID 7854253. Retrieved 27 January 2013.
{{cite journal}}: CS1 maint: date and year (link) - ^ Wakasugi, T.; Sugita, M.; Tsudzuki, T.; Sugiura, M. (1998). "Updated Gene Map of Tobacco Chloroplast DNA". Plant Molecular Biology Reporter. 16 (3): 231. doi:10.1023/A:1007564209282. S2CID 40036883. Retrieved 2 January 2013.
- ^ Krause K (September 2008). "From chloroplasts to "cryptic" plastids: evolution of plastid genomes in parasitic plants". Curr. Genet. 54 (3): 111–21. doi:10.1007/s00294-008-0208-8. PMID 18696071. S2CID 24879257.
{{cite journal}}: CS1 maint: date and year (link) - ^ Peng, Lianwei; Fukao, Yoichiro; Fujiwara, Masayuki; Shikanai, Toshiharu (24). "Multistep Assembly of Chloroplast NADH Dehydrogenase-Like Subcomplex A Requires Several Nucleus-Encoded Proteins, Including CRR41 and CRR42, in Arabidopsis". Plant Cell. 24 (1): 202–214. doi:10.1105/tpc.111.090597. PMC 3289569. PMID 22274627.
{{cite journal}}: Check date values in:|date=and|year=/|date=mismatch (help); Unknown parameter|month=ignored (help) - ^ Martin W, Rujan T, Richly E (September 2002). "Evolutionary analysis of Arabidopsis, cyanobacterial, and chloroplast genomes reveals plastid phylogeny and thousands of cyanobacterial genes in the nucleus". Proc. Natl. Acad. Sci. U.S.A. 99 (19): 12246–51. doi:10.1073/pnas.182432999. PMC 129430. PMID 12218172.
{{cite journal}}: CS1 maint: date and year (link) CS1 maint: multiple names: authors list (link) - ^ Huang CY, Ayliffe MA, Timmis JN (March 2003). "Direct measurement of the transfer rate of chloroplast DNA into the nucleus". Nature. 422 (6927): 72–6. doi:10.1038/nature01435. PMID 12594458. S2CID 4319507.
{{cite journal}}: CS1 maint: date and year (link) CS1 maint: multiple names: authors list (link) - ^ Koussevitzky, S.; Nott, A.; Mockler, T. C.; Hong, F.; Sachetto-Martins, G.; Surpin, M.; Lim, J.; Mittler, R.; Chory, J. (4). "Signals from Chloroplasts Converge to Regulate Nuclear Gene Expression". Science. 316 (5825): 715–719. doi:10.1126/science.+1140516 (inactive 29 July 2022). PMID 17395793. Retrieved 27 January 2013.
{{cite journal}}: Check date values in:|date=and|year=/|date=mismatch (help); Unknown parameter|month=ignored (help)CS1 maint: DOI inactive as of July 2022 (link) - ^ Hedtke, Boris; BöRner, Thomas; Weihe, Andreas (8). "Mitochondrial and Chloroplast Phage-Type RNA Polymerases in Arabidopsis". Science. 277 (5327): 809–811. doi:10.1126/science.277.5327.809. PMID 9242608. Retrieved 27 January 2013.
{{cite journal}}: Check date values in:|date=and|year=/|date=mismatch (help); Unknown parameter|month=ignored (help) - ^ Wise, R.R.; Hoober, J.K. (2007). The Structure and Function of Plastids. Springer. pp. 32–33. ISBN 9781402065705.
{{cite book}}: CS1 maint: multiple names: authors list (link) - ^ "Oedogonium Link ex Hirn, 1900: 17". algaeBASE. Retrieved 19 May 2013.
- ^ "Chlamydomonas Ehrenberg, 1833: 288". algaeBASE. Retrieved 19 May 2013.
- ^ "Spirogyra Link, 1820: 5". algaeBASE. Retrieved 19 May 2013.
- ^ "Sirogonium Kützing, 1843: 278". algaeBASE. Retrieved 19 May 2013.
- ^ "Zygnema C.Agardh, 1817: xxxii, 98". algaeBASE. Retrieved 19 May 2013.
- ^ "Micrasterias C.Agardh ex Ralfs, 1848: 68". algaeBASE. Retrieved 19 May 2013.
- ^ a b c d e f g Biology 8th edition—Campbell & Reece. Benjamin Cummings. 2008. pp. 196–197. ISBN 978-0-321-54325-7.
- ^ Koike, H.; Yoshio, M.; Kashino, Y.; Satoh, K. (1998 May). "Polypeptide composition of envelopes of spinach chloroplasts: two major proteins occupy 90% of outer envelope membranes" (PDF). Plant & Cell Physiology. 39 (5): 526–32. doi:10.1093/oxfordjournals.pcp.a029400. PMID 9664716. Retrieved 18 May 2013.
{{cite journal}}: Check date values in:|date=(help) - ^ a b Soll, Jürgen (March 2004). "Plant cell biology: Protein import into chloroplasts". Nature Reviews Molecular Cell Biology. 5 (3): 198–208. doi:10.1038/nrm1333. PMID 14991000. S2CID 32453554. Retrieved 19 May 2013.
{{cite journal}}: Unknown parameter|coauthors=ignored (|author=suggested) (help)CS1 maint: date and year (link) - ^ Köhler RH, Hanson MR (1 January 2000). "Plastid tubules of higher plants are tissue-specific and developmentally regulated". J. Cell. Sci. 113 (Pt 1): 81–9. doi:10.1242/jcs.113.1.81. PMID 10591627. Archived from the original on 18 November 2010.
- ^ Gray JC, Sullivan JA, Hibberd JM, Hansen MR (2001). "Stromules: mobile protrusions and interconnections between plastids". Plant Biology. 3 (3): 223–33. doi:10.1055/s-2001-15204.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ a b c d e f g h i j k Burgess, Jeremy (1989). An introduction to plant cell development ([Pbk. ed.). Cambridge: Cambridge university press. p. 46. ISBN 0521316111.
- ^ a b Whatley, Jean M (5). "The occurrence of a peripheral reticulum in plastids of the gymnosperm Welwitschia mirabilis". New Phytologist. 74 (2): 215–220. doi:10.1111/j.1469-8137.1975.tb02608.x. Retrieved 12 May 2013.
{{cite journal}}: Check date values in:|date=and|year=/|date=mismatch (help); Unknown parameter|month=ignored (help) - ^ a b c Wise, Robert R (2007). The Structure and Function of Plastids. Springer. pp. 17–18. ISBN 9781402065705.
- ^ Manuell, Andrea L.; Quispe, Joel; Mayfield, Stephen P. (1 January 2007). "Structure of the Chloroplast Ribosome: Novel Domains for Translation Regulation". PLOS Biology. 5 (8): e209. doi:10.1371/journal.pbio.0050209. PMC 1939882. PMID 17683199.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ a b c d Austin, Jotham R.; Frost, Elizabeth; Vidi, Pierre-Alexandre; Kessler, Felix; Staehelin, L. Andrew (July 2006). "Plastoglobules Are Lipoprotein Subcompartments of the Chloroplast That Are Permanently Coupled to Thylakoid Membranes and Contain Biosynthetic Enzymes". The Plant Cell. 18 (7): 1693–1703. doi:10.1105/tpc.105.039859. PMC 1488921. PMID 16731586. Retrieved 19 May 2013.
{{cite journal}}: CS1 maint: date and year (link) - ^ a b Crumpton-Taylor, Matilda; Grandison, Scott; Png, Kenneth M.Y.; Bushby, Andrew J.; Smith, Alison M. (December 2012). "Control of Starch Granule Numbers in Arabidopsis Chloroplasts". Plant Physiology. 158 (2): 905–916. doi:10.1104/pp.111.186957. PMC 3271777. PMID 22135430. Retrieved 19 May 2013.
{{cite journal}}: CS1 maint: date and year (link) - ^ Zeeman, Samuel C.; Delatte, Thierry; Messerli, Gaëlle; Umhang, Martin; Stettler, Michaela; Mettler, Tabea; Streb, Sebastian; Reinhold, Heike; Kötting, Oliver (1 January 2007). "Starch breakdown: recent discoveries suggest distinct pathways and novel mechanisms" (PDF). Functional Plant Biology. 34 (6): 465–473. doi:10.1071/FP06313. PMID 32689375. Retrieved 19 May 2013.
- ^ J D, Rochaix (1998). The molecular biology of chloroplasts and mitochondria in Chlamydomonas. Dordrecht [u.a.]: Kluwer Acad. Publ. pp. 550–565. ISBN 9780792351740.
- ^ a b c d e f Steer, Brian E.S. Gunning, Martin W. (1996). Plant cell biology : structure and function. Boston, Mass.: Jones and Bartlett Publishers. p. 24. ISBN 0867205040.
{{cite book}}: CS1 maint: multiple names: authors list (link) - ^ Hanson, David; Andrews, T. John; Badger, Murray R. (2002). "Variability of the pyrenoid-based CO2 concentrating mechanism in hornworts (Anthocerotophyta)". Functional Plant Biology. 29 (3): 407–416. doi:10.1071/PP01210. PMID 32689485. Retrieved 31 December 2012.
- ^ a b Ma, Yunbing; Pollock, Steve V.; Xiao, Ying; Cunnusamy, Khrishen; Moroney, James V. (2011). "Identification of a Novel Gene, CIA6, Required for Normal Pyrenoid Formation in Chlamydomonas reinhardtii". Plant Physiology. 156 (2): 884–96. doi:10.1104/pp.111.173922. PMC 3177283. PMID 21527423. Retrieved 31 December 2012.
- ^ a b Retallack, B.; Butler, R. D. (January 1970). "The Development and Structure of Pyrenoids in Bulbochaete hiloensis". Journal of Cell Science. 6 (1): 229–240. doi:10.1242/jcs.6.1.229. PMID 5417694. Retrieved 2 February 2013.
{{cite journal}}: CS1 maint: date and year (link) - ^ Brown, Malcolm R (1970). "Structure and Function of the Algal Pyrenoid" (PDF). Journal of Phycology. Retrieved 31 December 2012.
{{cite journal}}: Unknown parameter|coauthors=ignored (|author=suggested) (help) - ^ a b c d e Biology 8th edition—Campbell&Reece. Benjamin Cummings. 2008. pp. 186–187. ISBN 978-0-321-54325-7.
- ^ a b c d e f MustáRdy, LáSzló; Buttle, Karolyn; Steinbach, Gábor; Garab, Győző (24 October 2008). "The Three-Dimensional Network of the Thylakoid Membranes in Plants: Quasihelical Model of the Granum-Stroma Assembly". The Plant Cell. 20 (10): 2552–2557. doi:10.1105/tpc.108.059147. PMC 2590735. PMID 18952780. Retrieved 19 May 2013.
- ^ a b c Austin, Jotham R.; Staehelin, L. Andrew (2011). "Three-Dimensional Architecture of Grana and Stroma Thylakoids of Higher Plants as Determined by Electron Tomography". Plant Physiology. 155 (4): 1601–1611. doi:10.1104/pp.110.170647. PMC 3091084. PMID 21224341. Retrieved 18 May 2013.
- ^ "The Chloroplast Import Receptor Toc90 Partially Restores the Accumulation of Toc159 Client Proteins in the Arabidopsis thaliana ppi2 Mutant". Molecular Plant. Retrieved 19 May 2013.
- ^ "thylakoid". Merriam-Webster Dictionary. Merriam-Webster. Retrieved 19 May 2013.
- ^ a b c Biology 8th Edition Campbell & Reece. Benjamin Cummings (Pearson). 2009. pp. 190–193.
- ^ "Australian scientists discover first new chlorophyll in 60 years". University of Sydney. 20 August 2010.
- ^ a b c Takaichi, Shinichi (15 June 2011). "Carotenoids in Algae: Distributions, Biosyntheses and Functions". Marine Drugs. 9 (12): 1101–1118. doi:10.3390/md9061101. PMC 3131562. PMID 21747749.
- ^ Shapley, Dan. "Why Do Leaves Change Color in Fall?". News Articles. The Daily Green. Retrieved 21 May 2013.
- ^ "Introduction to the Rhodophyta". University of California Museum of Paleontology. Retrieved 20 May 2013.
- ^ a b c d e f g Biology 8th Edition Campbell & Reece. Benjamin Cummings (Pearson). 2009. pp. 200–201.
- ^ Lawton, June R (March 1988). "Ultrastructure of Chloroplast Membranes in Leaves of Maize and Ryegrass as Revealed by Selective Staining Methods". New Phytologist. 108 (3): 277–283. doi:10.1111/j.1469-8137.1988.tb04163.x. JSTOR 2433294. PMID 33873933. Retrieved 12 May 2013.
{{cite journal}}: CS1 maint: date and year (link) - ^ Roberts, editor, Keith (2007). Handbook of plant science. Chichester, West Sussex, England: Wiley. p. 16. ISBN 978-0-470-05723-0.
{{cite book}}:|first=has generic name (help)CS1 maint: multiple names: authors list (link) - ^ Biology 8th edition—Campbell&Reece. Benjamin Cummings. 2008. p. 742. ISBN 978-0-321-54325-7.
- ^ "Guard Cell Photosynthesis". Plant Physiology Online. Sinauer. Retrieved 14 April 2013.
- ^ Villee, Claude A. Plants: Their Biology and Importance. pp. 17, 127.
- ^ a b Eckardt, Nancy A (December 2003). "Controlling Organelle Positioning: A Novel Chloroplast Movement Protein". The Plant Cell. 15 (12): 2755–2757. doi:10.1105/tpc.151210. S2CID 5034665. Retrieved 6 March 2013.
{{cite journal}}: CS1 maint: date and year (link) - ^ a b c d e f g h i j Glynn, Jonathan M.; Miyagishima, Shin-ya; Yoder, David W.; Osteryoung, Katherine W.; Vitha, Stanislav (1 May 2007). "Chloroplast Division". Traffic. 8 (5): 451–461. doi:10.1111/j.1600-0854.2007.00545.x. PMID 17451550. S2CID 2808844. Retrieved 2 February 2013.
- ^ a b Dong, X.-J.; Nagai, R.; Takagi, S. (1998). "Microfilaments Anchor Chloroplasts along the Outer Periclinal Wall in Vallisneria Epidermal Cells through Cooperation of PFR and Photosynthesis" (PDF). Plant Cell Physiology. 39 (12): 1299–1306. doi:10.1093/oxfordjournals.pcp.a029334. Retrieved 6 March 2013.
- ^ a b c Takagi, Shingo (NaN undefined NaN). "Actin-based photo-orientation movement of chloroplasts in plant cells". Journal of Experimental Biology (206): 1963–1969. Retrieved 6 March 2013.
{{cite journal}}: Check date values in:|date=and|year=/|date=mismatch (help); Unknown parameter|month=ignored (help) - ^ a b c d e f g Padmanabhan, Meenu S.; Dinesh-Kumar, S. P. (1 November 2010). "All Hands on Deck—The Role of Chloroplasts, Endoplasmic Reticulum, and the Nucleus in Driving Plant Innate Immunity". Molecular Plant-Microbe Interactions. 23 (11): 1368–1380. doi:10.1094/MPMI-05-10-0113. PMID 20923348. Retrieved 4 February 2013.
{{cite journal}}: Unknown parameter|month=ignored (help) - ^ Katsir, Leron; Chung, Hoo Sun; Koo, Abraham JK; Howe, Gregg A. (2008). "Jasmonate signaling: a conserved mechanism of hormone sensing". Curr Bio. 11 (4): 428–435. doi:10.1016/j.pbi.2008.05.004. PMC 2560989. PMID 18583180.
- ^ Schnurr, Judy A.; Shockey, Jay M.; De Boer, Gert-Jan; Browse, John A. (July 2002). "Fatty Acid Export from the Chloroplast. Molecular Characterization of a Major Plastidial Acyl-Coenzyme A Synthetase from Arabidopsis". Plant Physiology. 129 (4): 1700–1709. doi:10.1104/pp.003251. PMC 166758. PMID 12177483. Retrieved 4 February 2013.
{{cite journal}}: CS1 maint: date and year (link) - ^ a b c d Biology—Concepts and Connections. Pearson. 2009. pp. 108–118.
- ^ a b Clarke, Jeremy M. Berg; John L. Tymoczko; Lubert Stryer. Web content by Neil D. (2002). Biochemistry (5. ed., 4. print. ed.). New York, NY [u.a.]: W. H. Freeman. pp. Section 19.4. ISBN 0-7167-3051-0.
{{cite book}}: CS1 maint: multiple names: authors list (link) - ^ a b Clarke, Jeremy M. Berg; John L. Tymoczko; Lubert Stryer. Web content by Neil D. (2002). Biochemistry (5. ed., 4. print. ed.). New York, NY [u.a.]: W. H. Freeman. pp. Section 20.1. ISBN 0-7167-3051-0.
{{cite book}}: CS1 maint: multiple names: authors list (link) - ^ a b c Wample, Robert L.; Davis, Ronald W. (1983 September). "Effect of Flooding on Starch Accumulation in Chloroplasts of Sunflower (Helianthus annuus L.)". Plant Physiology. 73 (1): 195–198. doi:10.1104/pp.73.1.195. PMC 1066435. PMID 16663176.
{{cite journal}}: Check date values in:|date=(help) - ^ Carmi, A.; Shomer, I. (1979). "Starch Accumulation and Photosynthetic Activity in Primary Leaves of Bean (Phaseolus vulgaris L.)". Annals of Botany. 44 (4): 479–484. doi:10.1093/oxfordjournals.aob.a085756.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Berg JM (2002). Biochemistry (5th ed.). W H Freeman. pp. Section 19.4. Retrieved 30 October 2012.
{{cite book}}: Unknown parameter|coauthors=ignored (|author=suggested) (help) - ^ a b c Hauser, M.; Eichelmann, H.; Oja, V.; Heber, U.; Laisk, A. (July 1995). "Stimulation by Light of Rapid pH Regulation in the Chloroplast Stroma in Vivo as lndicated by CO, Solubilization in Leaves". Plant Physiology. 108 (3): 1059–1066. doi:10.1104/pp.108.3.1059. PMC 157457. PMID 12228527. Retrieved 2 February 2013.
{{cite journal}}: CS1 maint: date and year (link) - ^ a b Werdan, Karl; Heldt, Hans W.; Milovancev, Mirjana (1975). "The role of pH in the regulation of carbon fixation in the chloroplast stroma. Studies on CO2 fixation in the light and dark". Science Direct. 396 (2): 276–292. doi:10.1016/0005-2728(75)90041-9. PMID 239746. Retrieved 9 December 2012.
- ^ Joyard, Jacques; Ferro, Myriam; Masselon, Christophe; Seigneurin-Berny, Daphné; Salvi, Daniel; Garin, Jérôme; Rolland, Norbert (23). "Chloroplast proteomics highlights the subcellular compartmentation of lipid metabolism" (PDF). Science Direct. 49 (2): 128–158. doi:10.1016/j.plipres.2009.10.003. PMID 19879895. Retrieved 28 December 2012.
{{cite journal}}: Check date values in:|date=and|year=/|date=mismatch (help); Unknown parameter|month=ignored (help) - ^ a b c d Burgess, Jeremy (1989). An introduction to plant cell development ([Pbk. ed.). Cambridge: Cambridge university press. p. 56. ISBN 0521316111.
- ^ Steer, Brian E.S. Gunning, Martin W. (1996). Plant cell biology : structure and function. Boston, Mass.: Jones and Bartlett Publishers. p. 20. ISBN 0867205040.
{{cite book}}: CS1 maint: multiple names: authors list (link) - ^ a b c d e Burgess, Jeremy (1989). An introduction to plant cell development ([Pbk. ed.). Cambridge: Cambridge university press. pp. 54–55. ISBN 0521316111.
- ^ Burgess, Jeremy (1989). An introduction to plant cell development ([Pbk. ed.). Cambridge: Cambridge university press. p. 57. ISBN 0521316111.
- ^ a b Possingham, J. V. (18 May 1976). "Chloroplast Replication and Chloroplast DNA Synthesis in Spinach Leaves" (PDF). Proceedings of the Royal Society B: Biological Sciences. 193 (1112): 295–305. doi:10.1098/rspb.1976.0047. S2CID 2691108. Retrieved 2 February 2013.
{{cite journal}}: Unknown parameter|coauthors=ignored (|author=suggested) (help) - ^ Heinhorst, S.; Cannon, G.C. (January 1993). "DNA replication in chloroplasts". Journal of Cell Science. 104 (104): 1–9. doi:10.1242/jcs.104.1.1. Retrieved 3 February 2013.
{{cite journal}}: CS1 maint: date and year (link) - ^ a b c d e f g h Miyagishima, Shin-ya; Nishida, Keiji; Mori, Toshiyuki; Matsuzaki, Motomichi; Higashiyama, Tetsuya; Kuroiwa, Haruko; Kuroiwa, Tsuneyoshi (February 2003). "A Plant-Specific Dynamin-Related Protein Forms a Ring at the Chloroplast Division Site". The Plant Cell. 15 (3): 655–665. doi:10.1105/tpc.009373. PMC 150020. PMID 12615939. Retrieved 2 February 2013.
{{cite journal}}: CS1 maint: date and year (link) - ^ a b Hashimoto, Haruki; Possingham, John Victor (April 1989). "Effect of Light on the Chloroplast Division Cycle and DNA Synthesis in Cultured Leaf Discs of Spinach". Plant Physiology. 89 (4): 1178–1183. doi:10.1104/pp.89.4.1178. PMC 1055993. PMID 16666681.
{{cite journal}}: CS1 maint: date and year (link) - ^ a b c d Hansen, A. Katie; Escobar, Linda K.; Gilbert, Lawrence E.; Jansen, Robert K. (1 January 2007). "Paternal, maternal, and biparental inheritance of the chloroplast genome in Passiflora (Passifloraceae): implications for phylogenetic studies". American Journal of Botany. 94 (1): 42–46. doi:10.3732/ajb.94.1.42. PMID 21642206. Retrieved 21 April 2013.
- ^ Birky Jr, C William (December 1995). "Uniparental inheritance of mitochondrial and chloroplast genes: Mechanisms and evolution" (PDF). Proceedings of the National Academy of Science. 92 (25): 11331–11338. doi:10.1073/pnas.92.25.11331. PMC 40394. PMID 8524780. Retrieved 21 April 2013.
{{cite journal}}: CS1 maint: date and year (link) - ^ Powell W, Morgante M, McDevitt R, Vendramin GG, Rafalski JA (August 1995). "Polymorphic simple sequence repeat regions in chloroplast genomes: applications to the population genetics of pines". Proc. Natl. Acad. Sci. U.S.A. 92 (17): 7759–63. doi:10.1073/pnas.92.17.7759. PMC 41225. PMID 7644491.
In the pines, the chloroplast genome is transmitted through pollen
{{cite journal}}: CS1 maint: date and year (link) CS1 maint: multiple names: authors list (link) - ^ Stegemann S, Hartmann S, Ruf S, Bock R (July 2003). "High-frequency gene transfer from the chloroplast genome to the nucleus". Proc. Natl. Acad. Sci. U.S.A. 100 (15): 8828–33. doi:10.1073/pnas.1430924100. PMC 166398. PMID 12817081.
most angiosperm species inherit their chloroplasts maternally
{{cite journal}}: CS1 maint: date and year (link) CS1 maint: multiple names: authors list (link) - ^ a b Ruf S, Karcher D, Bock R (April 2007). "Determining the transgene containment level provided by chloroplast transformation". Proc. Natl. Acad. Sci. U.S.A. 104 (17): 6998–7002. doi:10.1073/pnas.0700008104. PMC 1849964. PMID 17420459.
{{cite journal}}: CS1 maint: date and year (link) CS1 maint: multiple names: authors list (link)
External links
[edit]- Chloroplast - Cell Centered Database
- http://www.ncbi.nlm.nih.gov/nuccore/7525012?report=graphChloroplasts and Photosynthesis: The Role of Light from Kimball's Biology Pages
- Clegg MT, Gaut BS, Learn GH, Morton BR (July 1994). "Rates and patterns of chloroplast DNA evolution". Proc. Natl. Acad. Sci. U.S.A. 91 (15): 6795–801. doi:10.1073/pnas.91.15.6795. PMC 44285. PMID 8041699.
{{cite journal}}: CS1 maint: date and year (link) CS1 maint: multiple names: authors list (link) - 3D structures of proteins associated with thylakoid membrane
- Co-Extra research on chloroplast transformation
- NCBI full chloroplast genome
Category:Organelles Category:Photosynthesis Category:Endosymbiotic events
















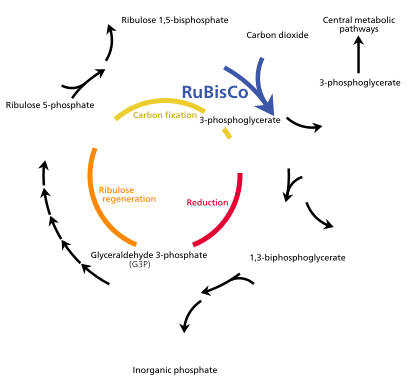
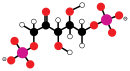




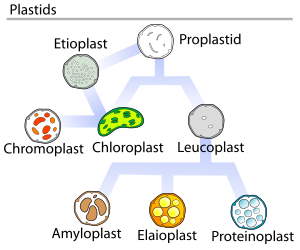

![Most chloroplasts in plant cells, and all chloroplasts in algae arise from chloroplast division.[109]](http://upload.wikimedia.org/wikipedia/commons/thumb/d/df/Chloroplast_division.svg/800px-Chloroplast_division.svg.png)