Reverse complement polymerase chain reaction
Reverse complement polymerase chain reaction (RC-PCR) is a modification of the polymerase chain reaction (PCR). It is primarily used to generate amplicon libraries for DNA sequencing by next generation sequencing (NGS). The technique permits both the amplification and the ability to append sequences or functional domains of choice independently to either end of the generated amplicons in a single closed tube reaction. RC-PCR was invented in 2013 by Daniel Ward and Christopher Mattocks at Salisbury NHS Foundation Trust, UK.
Principles
[edit]
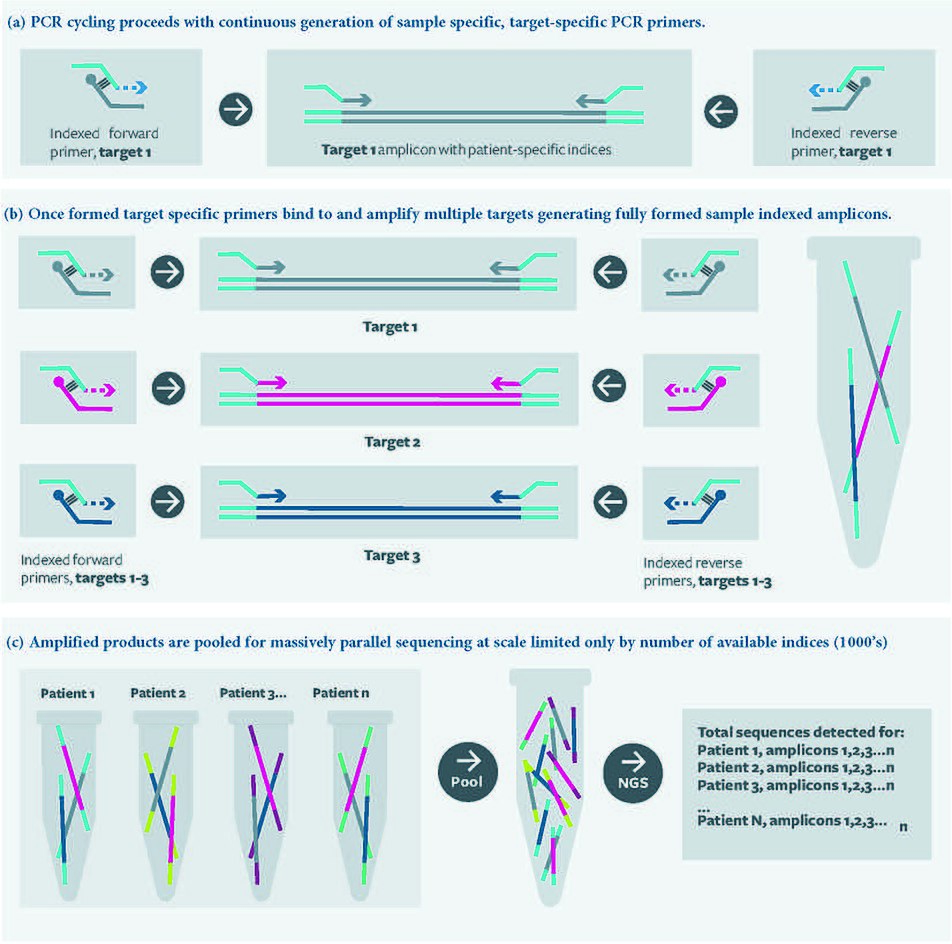
In RC-PCR, no target specific primers are present in the reaction mixture. Instead target specific primers are formed as the reaction proceeds. A typical reaction employing the approach requires four oligonucleotides. The oligonucleotides interact with each other in pairs; one oligonucleotide probe and one universal primer (containing functional domains of choice), which hybridize with each other at their 3’ ends. Once hybridized, the universal primer can be extended, using the oligonucleotide probe as the template, to yield fully formed, target specific primers, which are then available to amplify the template in subsequent rounds of thermal cycling as per a standard PCR reaction.
The oligonucleotide probe may also be blocked at the 3’ end preventing equivalent extension of the probe, but this is not essential. The probe is not consumed; it is available to act as a template for the universal primer to be ‘converted’ into target specific primer throughout successive PCR cycles. This generation of target specific primer occurs in parallel with standard PCR amplification under standard PCR conditions.
Advantages
[edit]
RC-PCR provides significant advantages over other methods of amplicon library preparation methods. Most significantly it is a single closed tube reaction, this eliminates cross contamination associated with other two-step PCR approaches as well as utilising less reagent and requiring less labour to perform.
The technique also provides the significant advantage of the flexibility of appending any desired sequence or functional domain of choice to either end of any amplicon. This is currently most advantageous in modern next generation sequencing (NGS) laboratories where a single target specific probe pair can be used with a whole library of universal primers. This benefit is used with NGS applications to apply sample specific indexes independently to each end of the amplicon construct. A Laboratory employing this approach would only require a single set of index primers, which can be used with all target specific probes compatible with that index set. This significantly reduces the number and length of oligonucleotides required by the laboratory compared to using full length pre-synthesised indexed target specific primers.
The generation of the target specific primer in the reaction as it progresses also leads to more balanced reaction components. Concentrations of target specific primer are more aligned with target molecule concentration thereby reducing the potential of both off target priming and primer dimerisation.
Variations
[edit]- Multiplex RC-PCR – where two or more universal primer probe sets are present in the reaction mixture to amplify two or more targets simultaneously.
- RT-RC-PCR – This modification is used when the template material supplied in the reaction is RNA rather than DNA. In this modification the reaction mixture also contains reverse transcriptase enzymes and reverse transcription primers as well as the universal primers and Reverse complement probes of the method. This approach permits reverse transcription of the provided RNA template, the formation of tailed target specific primers and the amplification of the desired targets in a single closed tube reaction.
- Single ended RC-PCR – This variation of the method is used when only one complementary universal primer probe pair is provided in the reaction to generate one target specific primer. The other target specific primer is provided as a traditional primer as per standard PCR.
History
[edit]Following the invention of RC-PCR in 2013 the technique was clinically validated and employed diagnostically for a range of both inherited diseases such as hemochromatosis and thrombophilia as well as somatically acquired disorders including Myeloproliferative neoplasms and Acute myeloid leukemia in the Wessex Regional Genetics Laboratory (WRGL), Salisbury UK. More recently work has been undertaken to utilise the technology in the fight against the SARS-CoV-2 pandemic.[1]
The patent application was filed in the UK in 2015 and awarded in 2020. Patent applications have been filed in other jurisdictions worldwide and are currently pending.
In May 2019 the Intellectual property was licensed to Nimagen B.V.[2] to develop, manufacture and market kits exploiting the technology. Currently commercially available kits employing the technology include those for Human identification[3][4] and more recently for the whole genome sequencing of the SARS-CoV-2 virus for variant identification, tracking and treatment response.[5][6] In August 2022 Nimagen officially launched a range of products employing the RC-PCR technology for human forensics applications under the trademark IDseek®.
The RC-PCR approach is becoming more widely used for human health and several CE IVD kits are available for human clinical diagnostics including BRCA, TP53, PALB2 and CFTR analysis. The technique has also been proven as a useful and powerful tool in the identification of the causative infectious pathogen in patients suspected of having a bacterial infection, in this setting it has been shown to provide a significant increase in the number of clinical samples in which a potentially clinically relevant pathogen is identified compared to the commonly used 16S Sanger method.[7] It has also been shown to provide similar advantages over traditional methods in the deconvolution of microbial communities in environmental samples.[8]
References
[edit]- ^ Mattocks, Christopher; Ward, Daniel; Mackay, Deborah (2021-03-05). "RT-RC-PCR: a novel and highly scalable next-generation sequencing method for simultaneous detection of SARS-COV-2 and typing variants of concern". medRxiv 10.1101/2021.03.02.21252704v1.
- ^ "NimaGen Licenses PCR Tech From Salisbury NHS Foundation Trust". Genomeweb. 2019-01-14. Retrieved 2021-04-22.
- ^ Kieser, Rachel E.; Buś, Magdalena M.; King, Jonathan L.; van der Vliet, Walter; Theelen, Joop; Budowle, Bruce (January 2020). "Reverse Complement PCR: A novel one-step PCR system for typing highly degraded DNA for human identification". Forensic Science International. Genetics. 44: 102201. doi:10.1016/j.fsigen.2019.102201. ISSN 1878-0326. PMID 31786458. S2CID 208535138.
- ^ Bus, Magdalena M; de Jong, Erik AC; King, Jonathan L; der Vliet, Walter van; Theelen, Joop; Budowle, Bruce (2021-08-05). "Reverse complement-PCR, an innovative and effective method for multiplexing forensically relevant single nucleotide polymorphism marker systems". BioTechniques. 71 (3): btn–2021–0031. doi:10.2144/btn-2021-0031. ISSN 0736-6205. PMID 34350776.
- ^ Wolters, Femke; Coolen, Jordy P. M.; Tostmann, Alma; Groningen, Lenneke F. J. van; Bleeker-Rovers, Chantal P.; Tan, Edward C. T. H.; Geest-Blankert, Nannet van der; Hautvast, Jeannine L. A.; Hopman, Joost; Wertheim, Heiman F. L.; Rahamat-Langendoen, Janette C. (2020-10-29). "Novel SARS-CoV-2 Whole-genome sequencing technique using Reverse Complement PCR enables fast and accurate outbreak analysis". bioRxiv: 2020.10.29.360578. doi:10.1101/2020.10.29.360578. S2CID 226228646.
- ^ Donovan-Banfield, I’ah; Penrice-Randal, Rebekah; Goldswain, Hannah; Rzeszutek, Aleksandra M.; Pilgrim, Jack; Bullock, Katie; Saunders, Geoffrey; Northey, Josh; Dong, Xiaofeng; Ryan, Yan; Reynolds, Helen; Tetlow, Michelle; Walker, Lauren E.; FitzGerald, Richard; Hale, Colin (2022-11-26). "Characterisation of SARS-CoV-2 genomic variation in response to molnupiravir treatment in the AGILE Phase IIa clinical trial". Nature Communications. 13 (1): 7284. Bibcode:2022NatCo..13.7284D. doi:10.1038/s41467-022-34839-9. ISSN 2041-1723. PMC 9701236. PMID 36435798.
- ^ Moorlag, Simone J. C. F. M.; Coolen, Jordy P. M.; van den Bosch, Bart; Jin, Elisabeth Hui-Mei; Buil, Jochem B.; Wertheim, Heiman F. L.; Melchers, Willem J. G. (2023-05-25). Luethy, Paul M. (ed.). "Targeting the 16S rRNA Gene by Reverse Complement PCR Next-Generation Sequencing: Specific and Sensitive Detection and Identification of Microbes Directly in Clinical Samples". Microbiology Spectrum. 11 (3). David Gaston: e0448322. doi:10.1128/spectrum.04483-22. hdl:2066/294286. ISSN 2165-0497. PMC 10269728. PMID 37227289.
- ^ Leontidou, Kleopatra; Abad-Recio, Ion L.; Rubel, Verena; Filker, Sabine; Däumer, Martin; Thielen, Alexander; Lanzén, Anders; Stoeck, Thorsten (2023-11-16). "Simultaneous analysis of seven <scp>16S rRNA</scp> hypervariable gene regions increases efficiency in marine bacterial diversity detection". Environmental Microbiology. 25 (12): 3484–3501. doi:10.1111/1462-2920.16530. ISSN 1462-2912. PMID 37974518.
