Prime editing
Prime editing is a 'search-and-replace' genome editing technology in molecular biology by which the genome of living organisms may be modified. The technology directly writes new genetic information into a targeted DNA site. It uses a fusion protein, consisting of a catalytically impaired Cas9 endonuclease fused to an engineered reverse transcriptase enzyme, and a prime editing guide RNA (pegRNA), capable of identifying the target site and providing the new genetic information to replace the target DNA nucleotides. It mediates targeted insertions, deletions, and base-to-base conversions without the need for double strand breaks (DSBs) or donor DNA templates.[1]
The technology has received mainstream press attention due to its potential uses in medical genetics. It utilizes methodologies similar to precursor genome editing technologies, including CRISPR/Cas9 and base editors. Prime editing has been used on some animal models of genetic disease[2][3][4] and plants.[5]
Genome editing
[edit]Components
[edit]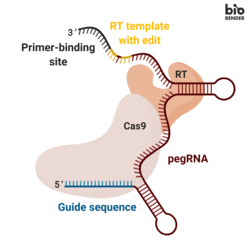
Prime editing involves three major components:[1]
- A prime editing guide RNA (pegRNA), capable of (i) identifying the target nucleotide sequence to be edited, and (ii) encoding new genetic information that replaces the targeted sequence. The pegRNA consists of an extended single guide RNA (sgRNA) containing a primer binding site (PBS) and a reverse transcriptase (RT) template sequence. During genome editing, the primer binding site allows the 3’ end of the nicked DNA strand to hybridize to the pegRNA, while the RT template serves as a template for the synthesis of edited genetic information.[1]
- A fusion protein consisting of a Cas9 H840A nickase fused to a Moloney Murine Leukemia Virus (M-MLV) reverse transcriptase.[1][6][7]
- Cas9 H840A nickase: the Cas9 enzyme contains two nuclease domains that can cleave DNA sequences, a RuvC domain that cleaves the non-target strand and a HNH domain that cleaves the target strand. The introduction of a H840A substitution in Cas9, through which the 840th amino acid histidine is replaced by an alanine, inactivates the HNH domain. With only the RuvC functioning domain, the catalytically impaired Cas9 introduces a single strand nick, hence the name nickase.[8]
- M-MLV reverse transcriptase: an enzyme that synthesizes DNA from a single-stranded RNA template.[1]
- A single guide RNA (sgRNA) that directs the Cas9 H840A nickase portion of the fusion protein to nick the non-edited DNA strand.[1]
Mechanism
[edit]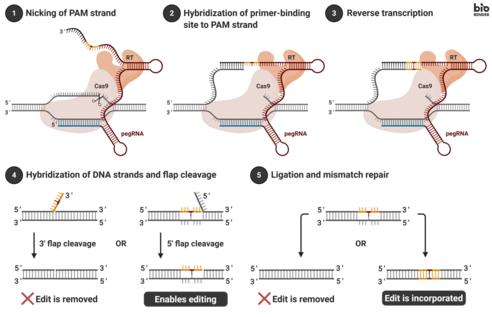
Genomic editing takes place by transfecting cells with the pegRNA and the fusion protein. Transfection is often accomplished by introducing vectors into a cell. Once internalized, the fusion protein nicks the target DNA sequence, exposing a 3’-hydroxyl group that can be used to initiate (prime) the reverse transcription of the RT template portion of the pegRNA. This results in a branched intermediate that contains two DNA flaps: a 3’ flap that contains the newly synthesized (edited) sequence, and a 5’ flap that contains the dispensable, unedited DNA sequence. The 5’ flap is then cleaved by structure-specific endonucleases or 5’ exonucleases. This process allows 3’ flap ligation, and creates a heteroduplex DNA composed of one edited strand and one unedited strand. The reannealed double stranded DNA contains nucleotide mismatches at the location where editing took place. In order to correct the mismatches, the cells exploit the intrinsic mismatch repair (MMR) mechanism, with two possible outcomes: (i) the information in the edited strand is copied into the complementary strand, permanently installing the edit; (ii) the original nucleotides are re-incorporated into the edited strand, excluding the edit.[1]
Development process
[edit]During the development of this technology, several modifications were done to the components, in order to increase its effectiveness.[1]
Prime editor 1
[edit]In the first system, a wild-type Moloney Murine Leukemia Virus (M-MLV) reverse transcriptase was fused to the Cas9 H840A nickase C-terminus. Detectable editing efficiencies were observed.[1]
Prime editor 2
[edit]In order to enhance DNA-RNA affinity, enzyme processivity, and thermostability, five amino acid substitutions were incorporated into the M-MLV reverse transcriptase. The mutant M-MLV RT was then incorporated into PE1 to give rise to (Cas9 (H840A)-M-MLV RT(D200N/L603W/T330P/T306K/W313F)). Efficiency improvement was observed over PE1.[1]

Prime editor 3
[edit]Despite its increased efficacy, the edit inserted by PE2 might still be removed due to DNA mismatch repair of the edited strand. To avoid this problem during DNA heteroduplex resolution, an additional single guide RNA (sgRNA) is introduced. This sgRNA is designed to match the edited sequence introduced by the pegRNA, but not the original allele. It directs the Cas9 nickase portion of the fusion protein to nick the unedited strand at a nearby site, opposite to the original nick. Nicking the non-edited strand causes the cell's natural repair system to copy the information in the edited strand to the complementary strand, permanently installing the edit.[1] However, there are drawbacks to this system as nicking the unaltered strand can lead to additional undesired indels. [9]
Prime editor 4
[edit]Prime editor 4 utilizes the same machinery as PE2, but also includes a plasmid that encodes for dominant negative MMR protein MLH1. Dominant negative MLH1 is able to essentially knock out endogenous MLH1 by inhibition, thereby reducing cellular MMR response and increasing prime editing efficiency.[9]
Prime editor 5
[edit]Prime editor 5 utilizes the same machinery as PE3, but also includes a plasmid that encodes for dominant negative MLH1. Like PE4, this allows for a knockdown of endogenous MMR response, increasing the efficiency of prime editing.[9]
Nuclease Prime Editor
[edit]Nuclease Prime Editor uses Cas9 nuclease instead of Cas9(H840A) nickase. Unlike prime editor 3 (PE3) that requires dual-nick at both DNA strands to induce efficient prime editing, Nuclease Prime Editor requires only a single pegRNA since the single-gRNA already creates double-strand break instead of single-strand nick.[10]
Twin prime editing
[edit]The "twin prime editing" (twinPE) mechanism reported in 2021 allows editing large sequences of DNA – sequences as large as genes – which addresses the method's key drawback. It uses a prime editor protein and two prime editing guide RNAs.[11][12][more detail needed]
History
[edit]Prime editing was developed in the lab of David R. Liu at the Broad Institute and disclosed in Anzalone et al. (2019).[13] Since then prime editing and the research that produced it have received widespread scientific acclaim,[14][6][15] being called "revolutionary"[7] and an important part of the future of editing.[13]
Development of epegRNAs
[edit]Prime editing efficiency can be increased with the use of engineered pegRNAs (epegRNAs). One common issue with traditional pegRNAs is degradation of the 3' end, leading to decreased PE efficiency. epegRNAs have a structured RNA motif added to their 3' end to prevent degradation.[16]
Implications
[edit]Although additional research is required to improve the efficiency of prime editing, the technology offers promising scientific improvements over other gene editing tools. The prime editing technology has the potential to correct the vast majority of pathogenic alleles that cause genetic diseases, as it can repair insertions, deletions, and nucleotide substitutions.[1]
Advantages
[edit]The prime editing tool offers advantages over traditional gene editing technologies. CRISPR/Cas9 edits rely on non-homologous end joining (NHEJ) or homology-directed repair (HDR) to fix DNA breaks, while the prime editing system employs DNA mismatch repair. This is an important feature of this technology given that DNA repair mechanisms such as NHEJ and HDR, generate unwanted, random insertions or deletions (INDELs). These are byproducts that complicate the retrieval of cells carrying the correct edit.[1][17]
The prime system introduces single-stranded DNA breaks instead of the double-stranded DNA breaks observed in other editing tools, such as base editors. Collectively, base editing and prime editing offer complementary strengths and weaknesses for making targeted transition mutations. Base editors offer higher editing efficiency and fewer INDEL byproducts if the desired edit is a transition point mutation and a PAM sequence exists roughly 15 bases from the target site. However, because the prime editing technology does not require a precisely positioned PAM sequence to target a nucleotide sequence, it offers more flexibility and editing precision. Remarkably, prime editors allow all types of substitutions, transitions and transversions to be inserted into the target sequence.[1][17] Cytosine base editing and adenine BE can already perform precise base transitions but for base transversions there have been no good options. Prime editing performs transversions with good usability. PE can insert up to 44bp, delete up to 80, or combinations thereof.[7]
Because the prime system involves three separate DNA binding events (between (i) the guide sequence and the target DNA, (ii) the primer binding site and the target DNA, and (iii) the 3’ end of the nicked DNA strand and the pegRNA), it has been suggested to have fewer undesirable off-target effects than CRISPR/Cas9.[1][17]
Limitations
[edit]There is considerable interest in applying gene-editing methods to the treatment of diseases with a genetic component. However, there are multiple challenges associated with this approach. An effective treatment would require editing of a large number of target cells, which in turn would require an effective method of delivery and a great level of tissue specificity.[1][18]
As of 2019, prime editing looks promising for relatively small genetic alterations, but more research needs to be conducted to evaluate whether the technology is efficient in making larger alterations, such as targeted insertions and deletions. Larger genetic alterations would require a longer RT template, which could hinder the efficient delivery of pegRNA to target cells. Furthermore, a pegRNA containing a long RT template could become vulnerable to damage caused by cellular enzymes.[1][18] Prime editing in plants suffers from low efficiency ranging from zero to a few percent and needs significant improvement.[19]
Some of these limitations have been mitigated by recent improvements to the prime editors,[2][20] including motifs that protect pegRNAs from degradation.[21] Further research is needed before prime editing could be used to correct pathogenic alleles in humans.[1][18] Research has also shown that inhibition of certain MMR proteins, including MLH1 can improve prime editing efficiency. [9]
Delivery method
[edit]Base editors used for prime editing require delivery of both a protein and RNA molecule into living cells. Introducing exogenous gene editing technologies into living organisms is a significant challenge. One potential way to introduce a base editor into animals and plants is to package the base editor into a viral capsid. The target organism can then be transduced by the virus to synthesize the base editor in vivo. Common laboratory vectors of transduction such as lentivirus cause immune responses in humans, so proposed human therapies often centered around adeno-associated virus (AAV) because AAV infections are largely asymptomatic. Unfortunately, the effective packaging capacity of AAV vectors is small, approximately 4.4kb not including inverted terminal repeats.[22] As a comparison, an SpCas9-reverse transcriptase fusion protein is 6.3kb,[1][23] which does not even account for the lengthened guide RNA necessary for targeting and priming the site of interest. However, successful delivery in mice has been achieved by splitting the editor into two AAV vectors[2][3][4][24] or by using an adenovirus,[3] which has a larger packaging capacity.
Applications
[edit]Prime editors may be used in gene drives. A prime editor may be incorporated into the Cleaver half of a Cleave and Rescue/ClvR system. In this case it is not meant to perform a precise alteration but instead to merely disrupt.[25]
PE is among recently introduced technologies which allow the transfer of single-nucleotide polymorphisms (SNPs) from one individual crop plant to another. PE is precise enough to be used to recreate an arbitrary SNP in an arbitrary target,[14] including deletions, insertions, and all 12 point mutations without also needing to perform a double-stranded break or carry a donating template.[6]
See also
[edit]References
[edit]- ^ a b c d e f g h i j k l m n o p q r s Anzalone, Andrew V.; Randolph, Peyton B.; Davis, Jessie R.; Sousa, Alexander A.; Koblan, Luke W.; Levy, Jonathan M.; Chen, Peter J.; Wilson, Christopher; Newby, Gregory A.; Raguram, Aditya; Liu, David R. (21 October 2019). "Search-and-replace genome editing without double-strand breaks or donor DNA". Nature. 576 (7785): 149–157. Bibcode:2019Natur.576..149A. doi:10.1038/s41586-019-1711-4. PMC 6907074. PMID 31634902.
- ^ a b c Liu, Pengpeng; Liang, Shun-Qing; Zheng, Chunwei; Mintzer, Esther; Zhao, Yan G.; Ponnienselvan, Karthikeyan; Mir, Aamir; Sontheimer, Erik J.; Gao, Guangping; Flotte, Terence R.; Wolfe, Scot A. (2021-04-09). "Improved prime editors enable pathogenic allele correction and cancer modelling in adult mice". Nature Communications. 12 (1): 2121. Bibcode:2021NatCo..12.2121L. doi:10.1038/s41467-021-22295-w. ISSN 2041-1723. PMC 8035190. PMID 33837189.
- ^ a b c Böck, Desirée; Rothgangl, Tanja; Villiger, Lukas; Schmidheini, Lukas; Mathis, Nicholas; Ioannidi, Eleonora; Kreutzer, Susanne; Kontarakis, Zacharias; Rimann, Nicole; Grisch-Chan, Hiu Man; Thöny, Beat (2021-08-17), Treatment of a metabolic liver disease by in vivo prime editing in mice, pp. 2021.08.17.456632, doi:10.1101/2021.08.17.456632, S2CID 237218057
- ^ a b Jang, Hyewon; Jo, Dong Hyun; Cho, Chang Sik; Shin, Jeong Hong; Seo, Jung Hwa; Yu, Goosang; Gopalappa, Ramu; Kim, Daesik; Cho, Sung-Rae; Kim, Jeong Hun; Kim, Hyongbum Henry (2021-08-26). "Application of prime editing to the correction of mutations and phenotypes in adult mice with liver and eye diseases". Nature Biomedical Engineering. 6 (2): 181–194. doi:10.1038/s41551-021-00788-9. ISSN 2157-846X. PMID 34446856. S2CID 237321703.
- ^ Biswas, Sudip; Bridgeland, Aya; Irum, Samra; Thomson, Michael J.; Septiningsih, Endang M. (29 August 2022). "Optimization of Prime Editing in Rice, Peanut, Chickpea, and Cowpea Protoplasts by Restoration of GFP Activity". International Journal of Molecular Sciences. 23 (17): 9809. doi:10.3390/ijms23179809. PMC 9456013. PMID 36077206.
- ^ a b c Lin, Xueqiu; Chemparathy, Augustine; La Russa, Marie; Daley, Timothy; Qi, Lei S. (2020-07-20). "Computational Methods for Analysis of Large-Scale CRISPR Screens". Annual Review of Biomedical Data Science. 3 (1). Annual Reviews: 137–162. doi:10.1146/annurev-biodatasci-020520-113523. ISSN 2574-3414. S2CID 225570135.
- ^ a b c Zhu, Haocheng; Li, Chao; Gao, Caixia (2020-09-24). "Applications of CRISPR–Cas in agriculture and plant biotechnology". Nature Reviews Molecular Cell Biology. 21 (11). Nature Portfolio: 661–677. doi:10.1038/s41580-020-00288-9. ISSN 1471-0072. PMID 32973356. S2CID 221918795.
- ^ Ran, F. Ann; Hsu, Patrick D.; Lin, Chie-Yu; Gootenberg, Jonathan S.; Konermann, Silvana; Trevino, Alexandro E.; Scott, David A.; Inoue, Azusa; Matoba, Shogo; Zhang, Yi; Zhang, Feng (September 2013). "Double Nicking by RNA-Guided CRISPR Cas9 for Enhanced Genome Editing Specificity". Cell. 154 (6): 1380–1389. doi:10.1016/j.cell.2013.08.021. PMC 3856256. PMID 23992846.
- ^ a b c d Chen, Peter (October 28, 2021). "Enhanced Prime Editing Systems By Manipulating Cellular Determinants of Editing Outcomes". Cell. 184 (22): 5635–5652.e29. doi:10.1016/j.cell.2021.09.018. PMC 8584034. PMID 34653350.
- ^ Adikusuma, Fatwa; Lushington, Caleb; Arudkumar, Jayshen; Godahewa, Gelshan; Chey, Yu C J; Gierus, Luke; Geiger, Ashleigh; Jain, Yatish; Reti, Daniel; Wilson, Laurence O W; Bower, Denis C; Thomas, Paul Q (17 September 2021). "Optimized nickase- and nuclease-based prime editing in human and mouse cells". Nucleic Acids Research. 49 (18): 10785–10795. doi:10.1093/nar/gkab792. PMC 8501948. PMID 34534334.
- ^ Dicorato, Allessandra. "New prime editing system inserts entire genes in human cells". Broad Institute of MIT. Retrieved 16 January 2022.
- ^ Anzalone, Andrew V.; Gao, Xin D.; Podracky, Christopher J.; Nelson, Andrew T.; Koblan, Luke W.; Raguram, Aditya; Levy, Jonathan M.; Mercer, Jaron A. M.; Liu, David R. (9 December 2021). "Programmable deletion, replacement, integration and inversion of large DNA sequences with twin prime editing". Nature Biotechnology. 40 (5): 731–740. doi:10.1038/s41587-021-01133-w. ISSN 1546-1696. PMC 9117393. PMID 34887556. S2CID 245012407.
- ^ a b Graff, Gregory D.; Sherkow, Jacob S. (2020-08-31). "Models of Technology Transfer for Genome-Editing Technologies". Annual Review of Genomics and Human Genetics. 21 (1). Annual Reviews: 509–534. doi:10.1146/annurev-genom-121119-100145. hdl:2142/110346. ISSN 1527-8204. PMID 32151165. S2CID 212652569.
- ^ a b Soyk, Sebastian; Benoit, Matthias; Lippman, Zachary B. (2020-11-23). "New Horizons for Dissecting Epistasis in Crop Quantitative Trait Variation". Annual Review of Genetics. 54 (1). Annual Reviews: 287–307. doi:10.1146/annurev-genet-050720-122916. ISSN 0066-4197. PMID 32870731. S2CID 221467135.
- ^ Faculty Opinions of Anzalone, Andrew V.; Randolph, Peyton B.; Davis, Jessie R.; Sousa, Alexander A.; Koblan, Luke W.; Levy, Jonathan M.; Chen, Peter J.; Wilson, Christopher; Newby, Gregory A.; Raguram, Aditya; Liu, David R. (2019). "Search-and-replace genome editing without double-strand breaks or donor DNA". Nature. 576 (7785): 149–157. Bibcode:2019Natur.576..149A. doi:10.1038/s41586-019-1711-4. PMC 6907074. PMID 31634902. Retrieved 2021-11-13.
- ^ Nelson, James (October 4, 2021). "Engineered pegRNAs improve prime editing efficiency". Nature Biotechnology. 40 (432): 402–410. doi:10.1038/s41587-021-01039-7. PMC 8930418. PMID 34608327.
- ^ a b c Sheridan, Cormac (7 November 2019). "Gene editing enters 'prime' time". Nature Biotechnology. doi:10.1038/d41587-019-00032-5. PMID 33154577. S2CID 209564966.
- ^ a b c "Scientist David Liu takes your questions on CRISPR and prime editing". STAT. 2019-11-06. Retrieved 2020-02-28.
- ^ Molla, Kutubuddin; Sretenovic, Simon; Bansal, Kailash C.; Qi, Yiping (13 September 2021). "Precise plant genome editing using base editors and prime editors". Nature Plants. 7 (9): 1166–1187. Bibcode:2021NatPl...7.1166M. doi:10.1038/s41477-021-00991-1. PMID 34518669. S2CID 237503774.
- ^ Chen, Peter J.; Hussmann, Jeffrey A.; Yan, Jun; Knipping, Friederike; Ravisankar, Purnima; Chen, Pin-Fang; Chen, Cidi; Nelson, James W.; Newby, Gregory A.; Sahin, Mustafa; Osborn, Mark J. (October 2021). "Enhanced prime editing systems by manipulating cellular determinants of editing outcomes". Cell. 184 (22): 5635–5652.e29. doi:10.1016/j.cell.2021.09.018. ISSN 0092-8674. PMC 8584034. PMID 34653350.
- ^ Nelson, James W.; Randolph, Peyton B.; Shen, Simon P.; Everette, Kelcee A.; Chen, Peter J.; Anzalone, Andrew V.; An, Meirui; Newby, Gregory A.; Chen, Jonathan C.; Hsu, Alvin; Liu, David R. (2021-10-04). "Engineered pegRNAs improve prime editing efficiency". Nature Biotechnology. 40 (3): 402–410. doi:10.1038/s41587-021-01039-7. ISSN 1546-1696. PMC 8930418. PMID 34608327. S2CID 238356160.
- ^ Wu, Zhijian; Yang, Hongyan; Colosi, Peter (2010). "Effect of Genome Size on AAV Vector Packaging". Molecular Therapy. 18 (1): 80–86. doi:10.1038/mt.2009.255. PMC 2839202. PMID 19904234.
- ^ "Addgene: PCMV-PE2".
- ^ Zhi, Shengyao; Chen, Yuxi; Wu, Guanglan; Wen, Jinkun; Wu, Jinni; Liu, Qianyi; Li, Yang; Kang, Rui; Hu, Sihui; Wang, Jiahui; Liang, Puping (July 2021). "Dual-AAV delivering split prime editor system for in vivo genome editing". Molecular Therapy. 30 (1): 283–294. doi:10.1016/j.ymthe.2021.07.011. ISSN 1525-0016. PMC 8753371. PMID 34298129. S2CID 236212353.
- ^ Hay, Bruce A.; Oberhofer, Georg; Guo, Ming (2021-01-07). "Engineering the Composition and Fate of Wild Populations with Gene Drive". Annual Review of Entomology. 66 (1). Annual Reviews: 407–434. doi:10.1146/annurev-ento-020117-043154. ISSN 0066-4170. PMID 33035437. S2CID 222257628.
