Internal loop
Internal-loops (also termed interior loops) in RNA are found where the double stranded RNA separates due to no Watson-Crick-Franklin base pairing between the nucleotides. Internal-loops differ from Stem-loops as they occur in middle of a stretch of double stranded RNA. The non-canonicoal residues result in the double helix becoming distorted due to unwinding, unstacking and kinking.
Internal-loops can be classified as either symmetrical or asymmetrical, with some asymmetrical internal-loops, also known as bulges. Many important structural motifs are composed of internal loops such as the C-loop,[1] the docking-elbow,[2] kink-turns (k-turn),[3][4] the right-angle,[5] the sarcin/ricin loops (also called bulged-G motifs),[6][7][8] the twist-up motif[9] and the UAA/GAN internal loop motif.[10]
See also
[edit]References
[edit]- ^ Lescoute, A; Leontis, NB; Massire, C; Westhof, E (2005). "Recurrent structural RNA motifs, Isostericity Matrices and sequence alignments". Nucleic Acids Research. 33 (8): 2395–409. doi:10.1093/nar/gki535. PMC 1087784. PMID 15860776.
- ^ Lehmann, J; Jossinet, F; Gautheret, D (May 1, 2013). "A universal RNA structural motif docking the elbow of tRNA in the ribosome, RNAse P and T-box leaders". Nucleic Acids Research. 41 (10): 5494–502. doi:10.1093/nar/gkt219. PMC 3664808. PMID 23580544.
- ^ Klein, D.J. (2001). "The kink-turn: a new RNA secondary structure motif". The EMBO Journal. 20 (15): 4214–4221. doi:10.1093/emboj/20.15.4214. ISSN 1460-2075. PMC 149158. PMID 11483524.
- ^ Schroeder, KT; McPhee, SA; Ouellet, J; Lilley, DM (Aug 2010). "A structural database for k-turn motifs in RNA". RNA. 16 (8): 1463–8. doi:10.1261/rna.2207910. PMC 2905746. PMID 20562215.
- ^ Grabow, WW; Zhuang, Z; Swank, ZN; Shea, JE; Jaeger, L (Nov 23, 2012). "The right angle (RA) motif: a prevalent ribosomal RNA structural pattern found in group I introns". Journal of Molecular Biology. 424 (1–2): 54–67. doi:10.1016/j.jmb.2012.09.012. PMC 3488136. PMID 22999957.
- ^ a b c Szewczak, AA; Moore, PB (Mar 17, 1995). "The sarcin/ricin loop, a modular RNA". Journal of Molecular Biology. 247 (1): 81–98. doi:10.1006/jmbi.1994.0124. PMID 7897662.
- ^ Leontis, NB; Westhof, E (Oct 30, 1998). "A common motif organizes the structure of multi-helix loops in 16 S and 23 S ribosomal RNAs". Journal of Molecular Biology. 283 (3): 571–83. doi:10.1006/jmbi.1998.2106. PMID 9784367.
- ^ Moore PB (1999). "Structural motifs in RNA". Annu. Rev. Biochem. 68: 287–300. doi:10.1146/annurev.biochem.68.1.287. PMID 10872451.
- ^ a b Zhong, C; Zhang, S (Feb 2012). "Clustering RNA structural motifs in ribosomal RNAs using secondary structural alignment". Nucleic Acids Research. 40 (3): 1307–17. doi:10.1093/nar/gkr804. PMC 3273805. PMID 21976732.
- ^ a b Lee, JC; Gutell, RR; Russell, R (Jul 28, 2006). "The UAA/GAN internal loop motif: a new RNA structural element that forms a cross-strand AAA stack and long-range tertiary interactions". Journal of Molecular Biology. 360 (5): 978–88. doi:10.1016/j.jmb.2006.05.066. PMID 16828489.


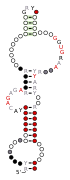



![A consensus secondary structure and primary sequence for the 3` Sarcin-Ricin (or bulged-G) RNA motif.[6]](http://upload.wikimedia.org/wikipedia/commons/thumb/f/f8/Sarcin-ricin-1-secondary-structure.svg/123px-Sarcin-ricin-1-secondary-structure.svg.png)
![A consensus secondary structure and primary sequence for the 5` Sarcin-Ricin (or bulged-G) RNA motif.[6]](http://upload.wikimedia.org/wikipedia/commons/thumb/7/73/Sarcin-ricin-2-secondary-structure.svg/152px-Sarcin-ricin-2-secondary-structure.svg.png)
![A consensus secondary structure and primary sequence for the twist-up RNA motif.[9]](http://upload.wikimedia.org/wikipedia/commons/thumb/9/98/Twist_up-secondary-structure.svg/100px-Twist_up-secondary-structure.svg.png)
![A consensus secondary structure and primary sequence for the UAA-GAN RNA motif.[10]](http://upload.wikimedia.org/wikipedia/commons/thumb/4/4e/UAA_GAN-secondary-structure.svg/74px-UAA_GAN-secondary-structure.svg.png)