Pseudomonas aeruginosa
| Pseudomonas aeruginosa | |
|---|---|

| |
| P. aeruginosa colonies on blood agar | |
| Scientific classification | |
| Domain: | Bacteria |
| Phylum: | Pseudomonadota |
| Class: | Gammaproteobacteria |
| Order: | Pseudomonadales |
| Family: | Pseudomonadaceae |
| Genus: | Pseudomonas |
| Species: | P. aeruginosa
|
| Binomial name | |
| Pseudomonas aeruginosa (Schröter 1872)
Migula 1900 | |
| Synonyms | |
| |

Pseudomonas aeruginosa is a common encapsulated, Gram-negative, aerobic–facultatively anaerobic, rod-shaped bacterium that can cause disease in plants and animals, including humans.[1][2] A species of considerable medical importance, P. aeruginosa is a multidrug resistant pathogen recognized for its ubiquity, its intrinsically advanced antibiotic resistance mechanisms, and its association with serious illnesses – hospital-acquired infections such as ventilator-associated pneumonia and various sepsis syndromes. P. aeruginosa is able to selectively inhibit various antibiotics from penetrating its outer membrane - and has high resistance to several antibiotics. According to the World Health Organization P. aeruginosa poses one of the greatest threats to humans in terms of antibiotic resistance.[3]
The organism is considered opportunistic insofar as serious infection often occurs during existing diseases or conditions – most notably cystic fibrosis and traumatic burns. It generally affects the immunocompromised but can also infect the immunocompetent as in hot tub folliculitis. Treatment of P. aeruginosa infections can be difficult due to its natural resistance to antibiotics. When more advanced antibiotic drug regimens are needed adverse effects may result.
It is citrate, catalase, and oxidase positive. It is found in soil, water, skin flora, and most human-made environments throughout the world. It thrives not only in normal atmospheres, but also in low-oxygen atmospheres, thus has colonized many natural and artificial environments. It uses a wide range of organic material for food; in animals, its versatility enables the organism to infect damaged tissues or those with reduced immunity. The symptoms of such infections are generalized inflammation and sepsis. If such colonizations occur in critical body organs, such as the lungs, the urinary tract, and kidneys, the results can be fatal.[4] Because it thrives on moist surfaces, this bacterium is also found on and in medical equipment, including catheters, causing cross-infections in hospitals and clinics. It is also able to decompose hydrocarbons and has been used to break down tarballs and oil from oil spills.[5] P. aeruginosa is not extremely virulent in comparison with other major species of pathogenic bacteria such as Gram-positive Staphylococcus aureus and Streptococcus pyogenes – though P. aeruginosa is capable of extensive colonization, and can aggregate into enduring biofilms.[6]
Nomenclature
[edit]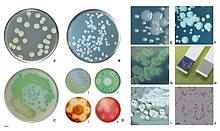
The word Pseudomonas means "false unit", from the Greek pseudēs (Greek: ψευδής, false) and (Latin: monas, from Greek: μονάς, a single unit). The stem word mon was used early in the history of microbiology to refer to microorganisms and germs, e.g., kingdom Monera.[7]
The species name aeruginosa is a Latin word meaning verdigris ("copper rust"), referring to the blue-green color of laboratory cultures of the species. This blue-green pigment is a combination of two secondary metabolites of P. aeruginosa, pyocyanin (blue) and pyoverdine (green), which impart the blue-green characteristic color of cultures.[7] Another assertion from 1956 is that aeruginosa may be derived from the Greek prefix ae- meaning "old or aged", and the suffix ruginosa means wrinkled or bumpy.[8]
The names pyocyanin and pyoverdine are from the Greek, with pyo-, meaning "pus",[9] cyanin, meaning "blue",[10] and verdine, meaning "green".[citation needed] Hence, the term "pyocyanic bacteria" refers specifically to the "blue pus" characteristic of a P. aeruginosa infection. Pyoverdine in the absence of pyocyanin is a fluorescent-yellow color.[citation needed]

Biology
[edit]Genome
[edit]The genome of Pseudomonas aeruginosa consists of a relatively large circular chromosome (5.5–6.8 Mb) that carries between 5,500 and 6,000 open reading frames, and sometimes plasmids of various sizes depending on the strain.[11] Comparison of 389 genomes from different P. aeruginosa strains showed that just 17.5% is shared. This part of the genome is the P. aeruginosa core genome.[12]
| strain: | VRFPA04 | C3719 | PAO1 | PA14 | PACS2 |
|---|---|---|---|---|---|
| Chromosome size (bp) | 6,818,030 | 6,222,097 | 6,264,404 | 6,537,648 | 6,492,423 |
| ORFs | 5,939 | 5,578 | 5,571 | 5,905 | 5,676 |

A comparative genomic study (in 2020) analyzed 494 complete genomes from the Pseudomonas genus, of which 189 were P. aeruginosa strains.[13] The study observed that their protein count and GC content ranged between 5500 and 7352 (average: 6192) and between 65.6 and 66.9% (average: 66.1%), respectively.[13] This comparative analysis further identified 1811 aeruginosa-core proteins, which accounts for more than 30% of the proteome. The higher percentage of aeruginosa-core proteins in this latter analysis could partly be attributed to the use of complete genomes. Although P. aeruginosa is a very well-defined monophyletic species, phylogenomically and in terms of ANIm values, it is surprisingly diverse in terms of protein content, thus revealing a very dynamic accessory proteome, in accordance with several analyses.[13][14][15][16] It appears that, on average, industrial strains have the largest genomes, followed by environmental strains, and then clinical isolates.[13][17] The same comparative study (494 Pseudomonas strains, of which 189 are P. aeruginosa) identified that 41 of the 1811 P. aeruginosa core proteins were present only in this species and not in any other member of the genus, with 26 (of the 41) being annotated as hypothetical. Furthermore, another 19 orthologous protein groups are present in at least 188/189 P. aeruginosa strains and absent in all the other strains of the genus.[citation needed]
Population structure
[edit]The population of P. aeruginosa can be classified in three main lineages, genetically characterised by the model strains PAO1, PA14, and the more divergent PA7.[18]
While P. aeruginosa is generally thought of as an opportunistic pathogen, several widespread clones appear to have become more specialised pathogens, particularly in cystic fibrosis patients, including the Liverpool epidemic strain (LES) which is found mainly in the UK,[19] DK2 in Denmark,[20] and AUST-02 in Australia (also previously known as AES-2 and P2).[21] There is also a clone that is frequently found infecting the reproductive tracts of horses.[22][23]
Metabolism
[edit]P. aeruginosa is a facultative anaerobe, as it is well adapted to proliferate in conditions of partial or total oxygen depletion. This organism can achieve anaerobic growth with nitrate or nitrite as a terminal electron acceptor. When oxygen, nitrate, and nitrite are absent, it is able to ferment arginine and pyruvate by substrate-level phosphorylation.[24] Additionally, phenazines produced by P. aeruginosa can act as electron shuttles to facilitate survival of cells at depth in biofilms.[25] Adaptation to microaerobic or anaerobic environments is essential for certain lifestyles of P. aeruginosa, for example, during lung infection in cystic fibrosis and primary ciliary dyskinesia, where thick layers of lung mucus and bacterially-produced alginate surrounding mucoid bacterial cells can limit the diffusion of oxygen. P. aeruginosa growth within the human body can be asymptomatic until the bacteria form a biofilm, which overwhelms the immune system. These biofilms are found in the lungs of people with cystic fibrosis and primary ciliary dyskinesia, and can prove fatal.[26][27][28][29][30][31][excessive citations]
Cellular co-operation
[edit]P. aeruginosa relies on iron as a nutrient source to grow. However, iron is not easily accessible because it is not commonly found in the environment. Iron is usually found in a largely insoluble ferric form.[32] Furthermore, excessively high levels of iron can be toxic to P. aeruginosa. To overcome this and regulate proper intake of iron, P. aeruginosa uses siderophores, which are secreted molecules that bind and transport iron.[33] These iron-siderophore complexes, however, are not specific. The bacterium that produced the siderophores does not necessarily receive the direct benefit of iron intake. Rather, all members of the cellular population are equally likely to access the iron-siderophore complexes. Members of the cellular population that can efficiently produce these siderophores are commonly referred to as cooperators; members that produce little to no siderophores are often referred to as cheaters. Research has shown when cooperators and cheaters are grown together, cooperators have a decrease in fitness, while cheaters have an increase in fitness.[34] The magnitude of change in fitness increases with increasing iron limitation.[35] With an increase in fitness, the cheaters can outcompete the cooperators; this leads to an overall decrease in fitness of the group, due to lack of sufficient siderophore production. These observations suggest that having a mix of cooperators and cheaters can reduce the virulent nature of P. aeruginosa.[34]
Enzymes
[edit]LigDs form a subfamily of the DNA ligases. These all have a LigDom/ligase domain, but many bacterial LigDs also have separate polymerase domains/PolDoms and nuclease domains/NucDoms. In P. aeruginosa's case the nuclease domains are N-terminus, and the polymerase domains are C-terminus, extensions of the single central ligase domain.[36]
Pathogenesis
[edit]This section needs additional citations for verification. (February 2021) |

Frequently acting as an opportunistic, nosocomial pathogen of immunocompromised individuals, but capable of infecting the immunocompetent, P. aeruginosa typically infects the airway, urinary tract, burns, and wounds, and also causes other blood infections.[37]
| Infections | Details and common associations | High-risk groups |
|---|---|---|
| Pneumonia | Diffuse bronchopneumonia | Cystic fibrosis, non-CF bronchiectasis patients |
| Septic shock | Associated with a purple-black skin lesion ecthyma gangrenosum | Neutropenic patients |
| Urinary tract infection | Urinary tract catheterization | |
| Gastrointestinal infection | Necrotising enterocolitis | Premature infants and neutropenic cancer patients |
| Skin and soft tissue infections | Hemorrhage and necrosis | People with burns or wound infections |
It is the most common cause of infections of burn injuries and of the outer ear (otitis externa), and is the most frequent colonizer of medical devices (e.g., catheters). Pseudomonas can be spread by equipment that gets contaminated and is not properly cleaned or on the hands of healthcare workers.[38] Pseudomonas can, in rare circumstances, cause community-acquired pneumonias,[39] as well as ventilator-associated pneumonias, being one of the most common agents isolated in several studies.[40] Pyocyanin is a virulence factor of the bacteria and has been known to cause death in C. elegans by oxidative stress. However, salicylic acid can inhibit pyocyanin production.[41] One in ten hospital-acquired infections is from Pseudomonas [citation needed]. Cystic fibrosis patients are also predisposed to P. aeruginosa infection of the lungs due to a functional loss in chloride ion movement across cell membranes as a result of a mutation.[42] P. aeruginosa may also be a common cause of "hot-tub rash" (dermatitis), caused by lack of proper, periodic attention to water quality. Since these bacteria thrive in moist environments, such as hot tubs and swimming pools, they can cause skin rash or swimmer's ear.[38] Pseudomonas is also a common cause of postoperative infection in radial keratotomy surgery patients. The organism is also associated with the skin lesion ecthyma gangrenosum. P. aeruginosa is frequently associated with osteomyelitis involving puncture wounds of the foot, believed to result from direct inoculation with P. aeruginosa via the foam padding found in tennis shoes, with diabetic patients at a higher risk.
A comparative genomic analysis of 494 complete Pseudomonas genomes, including 189 complete P. aeruginosa genomes, identified several proteins that are shared by the vast majority of P. aeruginosa strains, but are not observed in other analyzed Pseudomonas genomes.[13] These aeruginosa-specific core proteins, such as CntL, CntM, PlcB, Acp1, MucE, SrfA, Tse1, Tsi2, Tse3, and EsrC are known to play an important role in this species' pathogenicity.[13]
Toxins
[edit]P. aeruginosa uses the virulence factor exotoxin A to inactivate eukaryotic elongation factor 2 via ADP-ribosylation in the host cell, much as the diphtheria toxin does. Without elongation factor 2, eukaryotic cells cannot synthesize proteins and necrotise. The release of intracellular contents induces an immunologic response in immunocompetent patients. In addition P. aeruginosa uses an exoenzyme, ExoU, which degrades the plasma membrane of eukaryotic cells, leading to lysis. Increasingly, it is becoming recognized that the iron-acquiring siderophore, pyoverdine, also functions as a toxin by removing iron from mitochondria, inflicting damage on this organelle.[43][44] Since pyoverdine is secreted into the environment, it can be easily detected by the host or predator, resulting the host/predator migration towards the bacteria.[45]
Phenazines
[edit]Phenazines are redox-active pigments produced by P. aeruginosa. These pigments are involved in quorum sensing, virulence, and iron acquisition.[46] P. aeruginosa produces several pigments all produced by a biosynthetic pathway: phenazine-1-carboxamide (PCA), 1-hydroxyphenazine, 5-methylphenazine-1-carboxylic acid betaine, pyocyanin and aeruginosin A. Two nearly identical operons are involved in phenazine biosynthesis: phzA1B1C1D1E1F1G1 and phzA2B2C2D2E2F2G2.[47][48][49] The enzymes encoded by these operons convert chorismic acid to PCA. The products of three key genes, phzH, phzM, and phzS then convert PCA to the other phenazines mentioned above. Though phenazine biosynthesis is well studied, questions remain as to the final structure of the brown phenazine pyomelanin.[citation needed]
When pyocyanin biosynthesis is inhibited, a decrease in P. aeruginosa pathogenicity is observed in vitro. This suggests that pyocyanin is mostly responsible for the initial colonization of P. aeruginosa in vivo.[49]
Triggers
[edit]With low phosphate levels, P. aeruginosa has been found to activate from benign symbiont to express lethal toxins inside the intestinal tract and severely damage or kill the host, which can be mitigated by providing excess phosphate instead of antibiotics.[50]
Plants and invertebrates
[edit]In higher plants, P. aeruginosa induces soft rot, for example in Arabidopsis thaliana (Thale cress)[51] and Lactuca sativa (lettuce).[52][53] It is also pathogenic to invertebrate animals, including the nematode Caenorhabditis elegans,[54][55] the fruit fly Drosophila,[56] and the moth Galleria mellonella.[57] The associations of virulence factors are the same for plant and animal infections.[52][58] In both insects and plants, P. aeruginosa virulence is highly quorum sensing (QS) dependent.[59] Its QS is in turn highly dependent upon such genes as acyl-homoserine-lactone synthase, and lasI.[60]
Quorum sensing
[edit]P. aeruginosa is an opportunistic pathogen with the ability to coordinate gene expression in order to compete against other species for nutrients or colonization. Regulation of gene expression can occur through cell-cell communication or quorum sensing (QS) via the production of small molecules called autoinducers that are released into the external environment. These signals, when reaching specific concentrations correlated with specific population cell densities, activate their respective regulators thus altering gene expression and coordinating behavior. P. aeruginosa employs five interconnected QS systems – las, rhl, pqs, iqs, and pch – that each produce unique signaling molecules.[61] The las and rhl systems are responsible for the activation of numerous QS-controlled genes, the pqs system is involved in quinolone signaling, and the iqs system plays an important role in intercellular communication.[62] QS in P. aeruginosa is organized in a hierarchical manner. At the top of the signaling hierarchy is the las system, since the las regulator initiates the QS regulatory system by activating the transcription of a number of other regulators, such as rhl. So, the las system defines a hierarchical QS cascade from the las to the rhl regulons.[63] Detection of these molecules indicates P. aeruginosa is growing as biofilm within the lungs of cystic fibrosis patients.[64] The impact of QS and especially las systems on the pathogenicity of P. aeruginosa is unclear, however. Studies have shown that lasR-deficient mutants are associated with more severe outcomes in cystic fibrosis patients[65] and are found in up to 63% of chronically infected cystic fibrosis patients despite impaired QS activity.[66]
QS is known to control expression of a number of virulence factors in a hierarchical manner, including the pigment pyocyanin. However, although the las system initiates the regulation of gene expression, its absence does not lead to loss of virulence factors. Recently, it has been demonstrated that the rhl system partially controls las-specific factors, such as proteolytic enzymes responsible for elastolytic and staphylolytic activities, but in a delayed manner. So, las is a direct and indirect regulator of QS-controlled genes.[62] Another form of gene regulation that allows the bacteria to rapidly adapt to surrounding changes is through environmental signaling. Recent studies have discovered anaerobiosis can significantly impact the major regulatory circuit of QS. This important link between QS and anaerobiosis has a significant impact on production of virulence factors of this organism.[67] Garlic experimentally blocks quorum sensing in P. aeruginosa.[68]
Biofilms formation and cyclic di-GMP
[edit]As in most Gram negative bacteria, P. aeruginosa biofilm formation is regulated by one single molecule: cyclic di-GMP. At low cyclic di-GMP concentration, P. aeruginosa has a free-swimming mode of life. But when cyclic di-GMP levels increase, P. aeruginosa start to establish sessile communities on surfaces. The intracellular concentration of cyclic di-GMP increases within seconds when P. aeruginosa touches a surface (e.g.: a rock, plastic, host tissues...).[69] This activates the production of adhesive pili, that serve as "anchors" to stabilize the attachment of P. aeruginosa on the surface. At later stages, bacteria will start attaching irreversibly by producing a strongly adhesive matrix. At the same time, cyclic di-GMP represses the synthesis of the flagellar machinery, preventing P. aeruginosa from swimming. When suppressed, the biofilms are less adherent and easier to treat. The biofilm matrix of P. aeruginosa is composed of nucleic acids, amino acids, carbohydrates, and various ions. It mechanically and chemically protects P. aeruginosa from aggression by the immune system and some toxic compounds.[70] P. aeruginosa biofilm's matrix is composed of up to three types of sugar polymers (or "exopolysacharides") named PSL, PEL, and alginate.[71] Which exopolysacharides are produced varies by strain.[72]
- The polysaccharide synthesis operon and cyclic di-GMP form a positive feedback loop. This 15-gene operon is responsible for the cell-cell and cell-surface interactions required for cell communication.
- PEL is a cationic exopolysaccharide that cross-links extracellular DNA in the P. aeruginosa biofilm matrix.[73]
Upon certain cues or stresses, P. aeruginosa revert the biofilm program and detach. Recent studies have shown that the dispersed cells from P. aeruginosa biofilms have lower cyclic di-GMP levels and different physiologies from those of planktonic and biofilm cells,[74][75] with unique population dynamics and motility.[76] Such dispersed cells are found to be highly virulent against macrophages and C. elegans, but highly sensitive towards iron stress, as compared with planktonic cells.[74]
Biofilms and treatment resistance
[edit]Biofilms of P. aeruginosa can cause chronic opportunistic infections, which are a serious problem for medical care in industrialized societies, especially for immunocompromised patients and the elderly. They often cannot be treated effectively with traditional antibiotic therapy. Biofilms serve to protect these bacteria from adverse environmental factors, including host immune system components in addition to antibiotics. P. aeruginosa can cause nosocomial infections and is considered a model organism for the study of antibiotic-resistant bacteria. Researchers consider it important to learn more about the molecular mechanisms that cause the switch from planktonic growth to a biofilm phenotype and about the role of QS in treatment-resistant bacteria such as P. aeruginosa. This should contribute to better clinical management of chronically infected patients, and should lead to the development of new drugs.[67]
Scientists have been examining the possible genetic basis for P. aeruginosa resistance to antibiotics such as tobramycin. One locus identified as being an important genetic determinant of the resistance in this species is ndvB, which encodes periplasmic glucans that may interact with antibiotics and cause them to become sequestered into the periplasm. These results suggest a genetic basis exists behind bacterial antibiotic resistance, rather than the biofilm simply acting as a diffusion barrier to the antibiotic.[77]
Diagnosis
[edit]
Depending on the nature of infection, an appropriate specimen is collected and sent to a bacteriology laboratory for identification. As with most bacteriological specimens, a Gram stain is performed, which may show Gram-negative rods and/or white blood cells. P. aeruginosa produces colonies with a characteristic "grape-like" or "fresh-tortilla" odor on bacteriological media. In mixed cultures, it can be isolated as clear colonies on MacConkey agar (as it does not ferment lactose) which will test positive for oxidase. Confirmatory tests include production of the blue-green pigment pyocyanin on cetrimide agar and growth at 42 °C. A TSI slant is often used to distinguish nonfermenting Pseudomonas species from enteric pathogens in faecal specimens.[citation needed]
When P. aeruginosa is isolated from a normally sterile site (blood, bone, deep collections), it is generally considered dangerous, and almost always requires treatment.[78][79] However, P. aeruginosa is frequently isolated from nonsterile sites (mouth swabs, sputum, etc.), and, under these circumstances, it may represent colonization and not infection. The isolation of P. aeruginosa from nonsterile specimens should, therefore, be interpreted cautiously, and the advice of a microbiologist or infectious diseases physician/pharmacist should be sought prior to starting treatment. Often, no treatment is needed.[citation needed]
Classification
[edit]Morphological, physiological, and biochemical characteristics of Pseudomonas aeruginosa are shown in the Table below.
| Test type | Test | Characteristics |
|---|---|---|
| Colony characters | Size | Large |
| Type | Smooth | |
| Color | ||
| Shape | Flat | |
| Morphological characters | Shape | Rod |
| Physiological characters | Motility | + |
| Growth at 6.5% NaCl | - | |
| Biochemical characters | Gram staining | - |
| Oxidase | + | |
| Catalase | + | |
| Oxidative-Fermentative | ||
| Motility | + | |
| Methyl Red | - | |
| Voges-Proskauer | - | |
| Indole | - | |
| H2S Production | - | |
| Urease | - | |
| Nitrate reductase | + | |
| β-Galactosidase | ||
| Phenylalanine Deaminase | - | |
| DNAse | - | |
| Lipase | + | |
| Lysine Decarboxylase | - | |
| Pigment | + (bluish green pigmentation) | |
| Hemolysis | Beta/variable | |
| Hydrolysis of | Gelatin | + |
| Casein | ||
| Utilization of | Glycerol | + |
| Galactose | - | |
| D-Glucose | + | |
| D-Fructose | + | |
| D-Mannose | - | |
| Mannitol | + | |
| Citrate | + | |
| Maltose | - | |
| Sucrose | - | |
| Lactose | - |
Note: + = Positive, - =Negative
P. aeruginosa is a Gram-negative, aerobic (and at times facultatively anaerobic), rod-shaped bacterium with unipolar motility.[80] It has been identified as an opportunistic pathogen of both humans and plants.[81] P. aeruginosa is the type species of the genus Pseudomonas.[82]
Identification of P. aeruginosa can be complicated by the fact individual isolates often lack motility. The colony morphology itself also displays several varieties. The main two types are large, smooth, with a flat edge and elevated center and small, rough, and convex.[83] A third type, mucoid, can also be found. The large colony can typically be found in clinal settings while the small is found in nature.[83] The third, however, is present in biological settings and has been found in respiratory and in the urinary tract.[83] Furthermore, mutations in the gene lasR drastically alter colony morphology and typically lead to failure to hydrolyze gelatin or hemolyze.[citation needed]
In certain conditions, P. aeruginosa can secrete a variety of pigments, including pyocyanin (blue), pyoverdine (yellow and fluorescent), pyorubin (red), and pyomelanin (brown). These can be used to identify the organism.[84]

Clinical identification of P. aeruginosa may include identifying the production of both pyocyanin and fluorescein, as well as its ability to grow at 42 °C. P. aeruginosa is capable of growth in diesel and jet fuels, where it is known as a hydrocarbon-using microorganism, causing microbial corrosion.[85] It creates dark, gellish mats sometimes improperly called "algae" because of their appearance.[citation needed]
Treatment
[edit]This section needs additional citations for verification. (February 2021) |
Many P. aeruginosa isolates are resistant to a large range of antibiotics and may demonstrate additional resistance after unsuccessful treatment. It should usually be possible to guide treatment according to laboratory sensitivities, rather than choosing an antibiotic empirically. If antibiotics are started empirically, then every effort should be made to obtain cultures (before administering the first dose of antibiotic), and the choice of antibiotic used should be reviewed when the culture results are available.

Due to widespread resistance to many common first-line antibiotics, carbapenems, polymyxins, and more recently tigecycline were considered to be the drugs of choice; however, resistance to these drugs has also been reported. Despite this, they are still being used in areas where resistance has not yet been reported. Use of β-lactamase inhibitors such as sulbactam has been advised in combination with antibiotics to enhance antimicrobial action even in the presence of a certain level of resistance. Combination therapy after rigorous antimicrobial susceptibility testing has been found to be the best course of action in the treatment of multidrug-resistant P. aeruginosa. Some next-generation antibiotics that are reported as being active against P. aeruginosa include doripenem, ceftobiprole, and ceftaroline. However, these require more clinical trials for standardization. Therefore, research for the discovery of new antibiotics and drugs against P. aeruginosa is very much needed. Antibiotics that may have activity against P. aeruginosa include:
- aminoglycosides (gentamicin, amikacin, tobramycin, but not kanamycin)
- quinolones (ciprofloxacin, levofloxacin, but not moxifloxacin)
- cephalosporins (ceftazidime, cefepime, cefoperazone, cefpirome, ceftobiprole, but not cefuroxime, cefotaxime, or ceftriaxone)
- antipseudomonal penicillins: carboxypenicillins (carbenicillin and ticarcillin), and ureidopenicillins (mezlocillin, azlocillin, and piperacillin). P. aeruginosa is intrinsically resistant to all other penicillins.
- carbapenems (meropenem, imipenem, doripenem, but not ertapenem)
- polymyxins (polymyxin B and colistin)[86]
- monobactams (aztreonam)
As fluoroquinolones are one of the few antibiotic classes widely effective against P. aeruginosa, in some hospitals, their use is severely restricted to avoid the development of resistant strains. On the rare occasions where infection is superficial and limited (for example, ear infections or nail infections), topical gentamicin or colistin may be used[citation needed].
For pseudomonal wound infections, acetic acid with concentrations from 0.5% to 5% can be an effective bacteriostatic agent in eliminating the bacteria from the wound. Usually a sterile gauze soaked with acetic acid is placed on the wound after irrigation with normal saline. Dressing would be done once per day. Pseudomonas is usually eliminated in 90% of the cases after 10 to 14 days of treatment.[87]
Antibiotic resistance
[edit]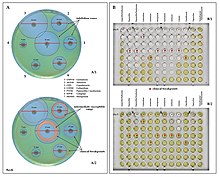
One of the most worrisome characteristics of P. aeruginosa is its low antibiotic susceptibility, which is attributable to a concerted action of multidrug efflux pumps with chromosomally encoded antibiotic resistance genes, i.e., the genes that encode proteins that serve as enzymes to break down antibiotics. Examples of such genes are:
- AmpC: encodes an AmpC-type β-lactamase enzyme, which breaks down penicillins, cephalosporins,[88] and carbapenems;[89]
- PER-1: encodes a PER-1 type extended-spectrum β-lactamase enzyme, which breaks down penicillins and cephalosporins;[90][91][92]
- IMP: encodes active-on-imipenem (IMP) carbapenemase (metallo-β-lactamase) enzyme which breaks down carbapenems;[93][94]
- NDM-1:[95] encodes a New Delhi metallo-β-lactamase 1 enzyme, which breaks down carbapenems;[96][97]
- OXA: encodes an oxacillinase (OCA) β-lactamase enzyme, which breaks down carbapenems;[98]
- AAC(6')-Ib: encodes an aminoglycoside-modifying enzyme called aminoglycoside N6'-acetyltransferase, which alters the structure of aminoglycoside antibiotics such as gentamicin and tobramycin;[99]
- Qnr: encodes a Qnr protein, which protects DNA gyrase and topoisomerase IV from the effects of quinolone (fluoroquinolone) antibiotics such as ciprofloxacin.[100]
Specific genes and enzymes involved in antibiotic resistance can vary between different strains.[101][102] P. aeruginosa TG523 harbored genes predicted to have antibacterial activity and those which are implicated in virulence.[103]
Another feature that contributes to antibiotic resistance of P. aeruginosa is the low permeability of the bacterial cellular envelopes.[104] In addition to this intrinsic resistance, P. aeruginosa easily develops acquired resistance either by mutation in chromosomally encoded genes or by the horizontal gene transfer of antibiotic resistance determinants. Development of multidrug resistance by P. aeruginosa isolates requires several different genetic events, including acquisition of different mutations and/or horizontal transfer of antibiotic resistance genes. Hypermutation favours the selection of mutation-driven antibiotic resistance in P. aeruginosa strains producing chronic infections, whereas the clustering of several different antibiotic resistance genes in integrons favors the concerted acquisition of antibiotic resistance determinants. Some recent studies have shown phenotypic resistance associated to biofilm formation or to the emergence of small-colony variants may be important in the response of P. aeruginosa populations to antibiotic treatment.[67]
Mechanisms underlying antibiotic resistance have been found to include production of antibiotic-degrading or antibiotic-inactivating enzymes, outer membrane proteins to evict the antibiotics, and mutations to change antibiotic targets. Presence of antibiotic-degrading enzymes such as extended-spectrum β-lactamases like PER-1, PER-2, and VEB-1, AmpC cephalosporinases, carbapenemases like serine oxacillinases, metallo-b-lactamases, OXA-type carbapenemases, and aminoglycoside-modifying enzymes, among others, have been reported. P. aeruginosa can also modify the targets of antibiotic action: for example, methylation of 16S rRNA to prevent aminoglycoside binding and modification of DNA, or topoisomerase to protect it from the action of quinolones. P. aeruginosa has also been reported to possess multidrug efflux pumps systems that confer resistance against a number of antibiotic classes, and the MexAB-OprM (Resistance-nodulation-division (RND) family) is considered as the most important[105]. An important factor found to be associated with antibiotic resistance is the decrease in the virulence capabilities of the resistant strain. Such findings have been reported in the case of rifampicin-resistant and colistin-resistant strains, in which decrease in infective ability, quorum sensing, and motility have been documented.[106]
Mutations in DNA gyrase are commonly associated with antibiotic resistance in P. aeruginosa. These mutations, when combined with others, confer high resistance without hindering survival. Additionally, genes involved in cyclic-di-GMP signaling may contribute to resistance. When P. aeruginosa is grown under in vitro conditions designed to mimic a cystic fibrosis patient's lungs, these genes mutate repeatedly.[107]
Two small RNAs, Sr0161 and ErsA, were shown to interact with mRNA encoding the major porin OprD responsible for the uptake of carbapenem antibiotics into the periplasm. The sRNAs bind to the 5'UTR of oprD, causing increase in bacterial resistance to meropenem. Another sRNA, Sr006, may positively regulate (post-transcriptionally) the expression of PagL, an enzyme responsible for deacylation of lipid A. This reduces the pro-inflammatory property of lipid A.[108] Furthermore, similar to a process found in Salmonella,[109] Sr006 regulation of PagL expression may aid in polymyxin B resistance.[108]
Prevention
[edit]Probiotic prophylaxis may prevent colonization and delay onset of Pseudomonas infection in an ICU setting.[110][non-primary source needed] Immunoprophylaxis against Pseudomonas is being investigated.[111] The risk of contracting P. aeruginosa can be reduced by avoiding pools, hot tubs, and other bodies of standing water; regularly disinfecting and/or replacing equipment that regularly encounters moisture (such as contact lens equipment and solutions); and washing one's hands often (which is protective against many other pathogens as well). However, even the best hygiene practices cannot totally protect an individual against P. aeruginosa, given how common P. aeruginosa is in the environment.[112]
Experimental therapies
[edit]Phage therapy against P. aeruginosa has been investigated as a possible effective treatment, which can be combined with antibiotics, has no contraindications and minimal adverse effects. Phages are produced as sterile liquid, suitable for intake, applications etc.[113] Phage therapy against ear infections caused by P. aeruginosa was reported in the journal Clinical Otolaryngology in August 2009.[114] As of 2024,[update] research on the topic is ongoing.[115]
Research
[edit]In 2013, João Xavier described an experiment in which P. aeruginosa, when subjected to repeated rounds of conditions in which it needed to swarm to acquire food, developed the ability to "hyperswarm" at speeds 25% faster than baseline organisms, by developing multiple flagella, whereas the baseline organism has a single flagellum.[116] This result was notable in the field of experimental evolution in that it was highly repeatable.[117] P. aeruginosa has been studied for use in bioremediation and use in processing polyethylene in municipal solid waste.[118]
Research on this bacterium's systems biology led to the development of genome-scale metabolic models that enable computer simulation and prediction of bacterial growth rates under varying conditions, including its virulence properties.[119][120]
Distribution
[edit]Pest risk analysis
[edit]As of 2019[update] the East African Community considers P. aeruginosa to be a quarantine concern because of the presence of Phaseolus vulgaris–pathogenic strains of P. aeruginosa in Kenya for the rest of the area. A pest risk analysis by the EAC was based on this bacterium's CABI's Crop Protection Compendium listing, following Kaaya & Darji 1989's initial detection in Kenya.[121]
Eyedrops
[edit]A small number of infections in the United States in 2022 and 2023 were likely caused by poorly manufactured eyedrops.[122]
See also
[edit]References
[edit]- ^ "UK Standards for Microbiology Investigations: Identification of Pseudomonas species and other Non- Glucose Fermenters" (PDF). Public Health England. 13 April 2015. Archived (PDF) from the original on 3 July 2022. Retrieved 4 May 2022.
- ^ Diggle SP, Whiteley M (January 2020). "Microbe Profile: Pseudomonas aeruginosa: opportunistic pathogen and lab rat". Microbiology. 166 (1): 30–33. doi:10.1099/mic.0.000860. PMC 7273324. PMID 31597590.
- ^ Spagnolo AM, Sartini M, Cristina ML (July 2021). "Pseudomonas aeruginosa in the healthcare facility setting". Reviews and Research in Medical Microbiology. 32 (3): 169–175. doi:10.1097/MRM.0000000000000271. ISSN 2770-3150. Archived from the original on 2024-01-18. Retrieved 2024-01-18.
- ^ Balcht A, Smith R (1994). Pseudomonas aeruginosa: Infections and Treatment. Informa Health Care. pp. 83–84. ISBN 978-0-8247-9210-7.
- ^ Itah AY, Essien JP (2005). "Growth Profile and Hydrocarbonoclastic Potential of Microorganisms Isolated from Tarballs in the Bight of Bonny, Nigeria". World Journal of Microbiology and Biotechnology. 21 (6–7): 1317–22. doi:10.1007/s11274-004-6694-z. S2CID 84888286.
- ^ Høiby N, Ciofu O, Bjarnsholt T (November 2010). "Pseudomonas aeruginosa biofilms in cystic fibrosis". Future Microbiology. 5 (11): 1663–1674. doi:10.2217/fmb.10.125. PMID 21133688.
- ^ a b Palleroni NJ (June 2010). "The Pseudomonas story". Environmental Microbiology. 12 (6): 1377–1383. doi:10.3201/eid1808.ET1808. PMC 3423701. PMID 20553550.
- ^ Brown RW (1956). Composition of Scientific Words. Smithsonian Institutional Press. ISBN 978-0-87474-286-2.
- ^ Tzouchas A (2014). WestBow Press. Greek Words. p. 550. ISBN 978-1-4907-2610-6. Archived from the original on 2023-04-26. Retrieved 2020-11-02.
- ^ Gonçalves T, Vasconcelos U (2021). "Colour Me Blue: The History and the Biotechnological Potential of Pyocyanin". Molecules. 26 (4): 927. doi:10.3390/molecules26040927. PMC 7916356. PMID 33578646.
- ^ Klockgether J, Cramer N, Wiehlmann L, Davenport CF, Tümmler B (2011). "Pseudomonas aeruginosa Genomic Structure and Diversity". Frontiers in Microbiology. 2: 150. doi:10.3389/fmicb.2011.00150. PMC 3139241. PMID 21808635.
- ^ De Smet J, Hendrix H, Blasdel BG, Danis-Wlodarczyk K, Lavigne R (September 2017). "Pseudomonas predators: understanding and exploiting phage-host interactions". Nature Reviews. Microbiology. 15 (9): 517–530. doi:10.1038/nrmicro.2017.61. PMID 28649138. S2CID 826136.
- ^ a b c d e f Nikolaidis M, Mossialos D, Oliver SG, Amoutzias GD (2020-07-24). "Comparative Analysis of the Core Proteomes among the Pseudomonas Major Evolutionary Groups Reveals Species-Specific Adaptations for Pseudomonas aeruginosa and Pseudomonas chlororaphis". Diversity. 12 (8): 289. doi:10.3390/d12080289. ISSN 1424-2818.
- ^ Ozer EA, Allen JP, Hauser AR (August 2014). "Characterization of the core and accessory genomes of Pseudomonas aeruginosa using bioinformatic tools Spine and AGEnt". BMC Genomics. 15 (1): 737. doi:10.1186/1471-2164-15-737. PMC 4155085. PMID 25168460.
- ^ Subedi D, Vijay AK, Kohli GS, Rice SA, Willcox M (October 2018). "Comparative genomics of clinical strains of Pseudomonas aeruginosa strains isolated from different geographic sites". Scientific Reports. 8 (1): 15668. Bibcode:2018NatSR...815668S. doi:10.1038/s41598-018-34020-7. PMC 6199293. PMID 30353070.
- ^ Freschi L, Vincent AT, Jeukens J, Emond-Rheault JG, Kukavica-Ibrulj I, Dupont MJ, et al. (January 2019). Martin B (ed.). "The Pseudomonas aeruginosa Pan-Genome Provides New Insights on Its Population Structure, Horizontal Gene Transfer, and Pathogenicity". Genome Biology and Evolution. 11 (1): 109–120. doi:10.1093/gbe/evy259. PMC 6328365. PMID 30496396.
- ^ Weiser R, Green AE, Bull MJ, Cunningham-Oakes E, Jolley KA, Maiden MC, et al. (July 2019). "Not all Pseudomonas aeruginosa are equal: strains from industrial sources possess uniquely large multireplicon genomes". Microbial Genomics. 5 (7). doi:10.1099/mgen.0.000276. PMC 6700666. PMID 31170060.
- ^ Roy PH, Tetu SG, Larouche A, Elbourne L, Tremblay S, Ren Q, et al. (January 2010). "Complete genome sequence of the multiresistant taxonomic outlier Pseudomonas aeruginosa PA7". PLOS ONE. 5 (1): e8842. Bibcode:2010PLoSO...5.8842R. doi:10.1371/journal.pone.0008842. PMC 2809737. PMID 20107499.
- ^ Winstanley C, Langille MG, Fothergill JL, Kukavica-Ibrulj I, Paradis-Bleau C, Sanschagrin F, et al. (January 2009). "Newly introduced genomic prophage islands are critical determinants of in vivo competitiveness in the Liverpool Epidemic Strain of Pseudomonas aeruginosa". Genome Research. 19 (1): 12–23. doi:10.1101/gr.086082.108. PMC 2612960. PMID 19047519.
- ^ Marvig RL, Johansen HK, Molin S, Jelsbak L (2013). "Genome analysis of a transmissible lineage of pseudomonas aeruginosa reveals pathoadaptive mutations and distinct evolutionary paths of hypermutators". PLOS Genetics. 9 (9): e1003741. doi:10.1371/journal.pgen.1003741. PMC 3764201. PMID 24039595.
- ^ Wee BA, Tai AS, Sherrard LJ, Ben Zakour NL, Hanks KR, Kidd TJ, et al. (August 2018). "Whole genome sequencing reveals the emergence of a Pseudomonas aeruginosa shared strain sub-lineage among patients treated within a single cystic fibrosis centre". BMC Genomics. 19 (1): 644. doi:10.1186/s12864-018-5018-x. PMC 6117919. PMID 30165811.
- ^ Kidd TJ, Ritchie SR, Ramsay KA, Grimwood K, Bell SC, Rainey PB (6 September 2012). "Pseudomonas aeruginosa exhibits frequent recombination, but only a limited association between genotype and ecological setting". PLOS ONE. 7 (9): e44199. Bibcode:2012PLoSO...744199K. doi:10.1371/journal.pone.0044199. PMC 3435406. PMID 22970178.
- ^ Kidd TJ, Gibson JS, Moss S, Greer RM, Cobbold RN, Wright JD, et al. (May 2011). "Clonal complex Pseudomonas aeruginosa in horses". Veterinary Microbiology. 149 (3–4): 508–512. doi:10.1016/j.vetmic.2010.11.030. PMID 21183294.
- ^ Schobert M, Jahn D (December 2010). "Anaerobic physiology of Pseudomonas aeruginosa in the cystic fibrosis lung". International Journal of Medical Microbiology. 300 (8): 549–556. doi:10.1016/j.ijmm.2010.08.007. PMID 20951638.
- ^ Dietrich LE, Okegbe C, Price-Whelan A, Sakhtah H, Hunter RC, Newman DK (April 2013). "Bacterial community morphogenesis is intimately linked to the intracellular redox state". Journal of Bacteriology. 195 (7): 1371–1380. doi:10.1128/JB.02273-12. PMC 3624522. PMID 23292774.
- ^ Tortora GJ, Case CL, Bair III WB, Weber D, Funke BR (2016). Microbiology: An Introduction (12th ed.). Pearson Education. p. 54. ISBN 978-0-321-92915-0.
- ^ Hassett DJ (December 1996). "Anaerobic production of alginate by Pseudomonas aeruginosa: alginate restricts diffusion of oxygen". Journal of Bacteriology. 178 (24): 7322–7325. doi:10.1128/jb.178.24.7322-7325.1996. PMC 178651. PMID 8955420.
- ^ Worlitzsch D, Tarran R, Ulrich M, Schwab U, Cekici A, Meyer KC, et al. (February 2002). "Effects of reduced mucus oxygen concentration in airway Pseudomonas infections of cystic fibrosis patients". The Journal of Clinical Investigation. 109 (3): 317–325. doi:10.1172/JCI13870. PMC 150856. PMID 11827991.
- ^ Cooper M, Tavankar GR, Williams HD (May 2003). "Regulation of expression of the cyanide-insensitive terminal oxidase in Pseudomonas aeruginosa". Microbiology. 149 (Pt 5): 1275–1284. doi:10.1099/mic.0.26017-0. PMID 12724389.
- ^ Williams HD, Zlosnik JE, Ryall B (2007). Oxygen, cyanide and energy generation in the cystic fibrosis pathogen Pseudomonas aeruginosa. Advances in Microbial Physiology. Vol. 52. pp. 1–71. doi:10.1016/S0065-2911(06)52001-6. ISBN 978-0-12-027752-0. PMID 17027370.
- ^ Leach R, Moore K, Bell D (2016). Oxford Desk Reference: Acute Medicine. Oxford University Press. p. 244. ISBN 978-0-19-100714-9.
- ^ Buckling A, Harrison F, Vos M, Brockhurst MA, Gardner A, West SA, et al. (November 2007). "Siderophore-mediated cooperation and virulence in Pseudomonas aeruginosa". FEMS Microbiology Ecology. 62 (2): 135–141. Bibcode:2007FEMME..62..135B. doi:10.1111/j.1574-6941.2007.00388.x. PMID 17919300.
- ^ Nguyen AT, Jones JW, Ruge MA, Kane MA, Oglesby-Sherrouse AG (July 2015). "Iron Depletion Enhances Production of Antimicrobials by Pseudomonas aeruginosa". Journal of Bacteriology. 197 (14): 2265–2275. doi:10.1128/JB.00072-15. PMC 4524187. PMID 25917911.
- ^ a b Harrison F, Browning LE, Vos M, Buckling A (July 2006). "Cooperation and virulence in acute Pseudomonas aeruginosa infections". BMC Biology. 4: 21. doi:10.1186/1741-7007-4-21. PMC 1526758. PMID 16827933.
- ^ Griffin AS, West SA, Buckling A (August 2004). "Cooperation and competition in pathogenic bacteria". Nature. 430 (7003): 1024–1027. Bibcode:2004Natur.430.1024G. doi:10.1038/nature02744. hdl:1842/698. PMID 15329720. S2CID 4429250.
- ^ Pitcher RS, Brissett NC, Doherty AJ (2007). "Nonhomologous end-joining in bacteria: a microbial perspective". Annual Review of Microbiology. 61 (1). Annual Reviews: 259–282. doi:10.1146/annurev.micro.61.080706.093354. PMID 17506672.
- ^ "Pseudomonas aeruginosa". Todar's Online Textbook of Bacteriology. 4 June 2004. Archived from the original on 9 October 2006. Retrieved 9 September 2011 – via Textbookofbacteriology.net.
- ^ a b "Pseudomonas aeruginosa in Healthcare Settings". Healthcare-associated Infections (HAI): Diseases and Organisms. Centers for Disease Control and Prevention. 7 May 2014. Archived from the original on 29 December 2017. Retrieved 8 September 2017.
- ^ Fine MJ, Smith MA, Carson CA, Mutha SS, Sankey SS, Weissfeld LA, et al. (January 1996). "Prognosis and outcomes of patients with community-acquired pneumonia. A meta-analysis". JAMA. 275 (2): 134–141. doi:10.1001/jama.1996.03530260048030. PMID 8531309.
- ^ Diekema DJ, Pfaller MA, Jones RN, Doern GV, Winokur PL, Gales AC, et al. (September 1999). "Survey of bloodstream infections due to gram-negative bacilli: frequency of occurrence and antimicrobial susceptibility of isolates collected in the United States, Canada, and Latin America for the SENTRY Antimicrobial Surveillance Program, 1997". Clinical Infectious Diseases. 29 (3): 595–607. doi:10.1086/598640. PMID 10530454.
- ^ Prithiviraj B, Bais HP, Weir T, Suresh B, Najarro EH, Dayakar BV, et al. (September 2005). "Down regulation of virulence factors of Pseudomonas aeruginosa by salicylic acid attenuates its virulence on Arabidopsis thaliana and Caenorhabditis elegans". Infection and Immunity. 73 (9): 5319–5328. doi:10.1128/IAI.73.9.5319-5328.2005. PMC 1231131. PMID 16113247.
- ^ Johnson PA (March 2019). "Novel understandings of host cell mechanisms involved in chronic lung infection: Pseudomonas aeruginosa in the cystic fibrotic lung". Journal of Infection and Public Health. 12 (2): 242–246. doi:10.1016/j.jiph.2018.10.014. PMID 30459101.
- ^ Kirienko NV, Ausubel FM, Ruvkun G (February 2015). "Mitophagy confers resistance to siderophore-mediated killing by Pseudomonas aeruginosa". Proceedings of the National Academy of Sciences of the United States of America. 112 (6): 1821–1826. Bibcode:2015PNAS..112.1821K. doi:10.1073/pnas.1424954112. PMC 4330731. PMID 25624506.
- ^ Kirienko NV, Kirienko DR, Larkins-Ford J, Wählby C, Ruvkun G, Ausubel FM (April 2013). "Pseudomonas aeruginosa disrupts Caenorhabditis elegans iron homeostasis, causing a hypoxic response and death". Cell Host & Microbe. 13 (4): 406–416. doi:10.1016/j.chom.2013.03.003. PMC 3641844. PMID 23601103.
- ^ Hu M, Ma Y, Chua SL (January 2024). "Bacterivorous nematodes decipher microbial iron siderophores as prey cue in predator-prey interactions". Proceedings of the National Academy of Sciences of the United States of America. 121 (3): e2314077121. Bibcode:2024PNAS..12114077H. doi:10.1073/pnas.2314077121. PMC 10801909. PMID 38190542.
- ^ Dietrich LE, Price-Whelan A, Petersen A, Whiteley M, Newman DK (September 2006). "The phenazine pyocyanin is a terminal signalling factor in the quorum sensing network of Pseudomonas aeruginosa". Molecular Microbiology. 61 (5): 1308–1321. doi:10.1111/j.1365-2958.2006.05306.x. PMID 16879411. S2CID 4985392.
- ^ Abu EA, Su S, Sallans L, Boissy RE, Greatens A, Heineman WR, et al. (August 2013). "Cyclic voltammetric, fluorescence and biological analysis of purified aeruginosin A, a secreted red pigment of Pseudomonas aeruginosa PAO1". Microbiology. 159 (Pt 8): 1736–1747. doi:10.1099/mic.0.065235-0. PMID 23782801.
- ^ Mavrodi DV, Bonsall RF, Delaney SM, Soule MJ, Phillips G, Thomashow LS (November 2001). "Functional analysis of genes for biosynthesis of pyocyanin and phenazine-1-carboxamide from Pseudomonas aeruginosa PAO1". Journal of Bacteriology. 183 (21): 6454–6465. doi:10.1128/JB.183.21.6454-6465.2001. PMC 100142. PMID 11591691.
- ^ a b Ho Sui SJ, Lo R, Fernandes AR, Caulfield MD, Lerman JA, Xie L, et al. (September 2012). "Raloxifene attenuates Pseudomonas aeruginosa pyocyanin production and virulence". International Journal of Antimicrobial Agents. 40 (3): 246–251. doi:10.1016/j.ijantimicag.2012.05.009. PMC 5511546. PMID 22819149.
- ^ "Research could lead to new non-antibiotic drugs to counter hospital infections" (Press release). University of Chicago Medical Center. 2009-04-14. Archived from the original on 2022-06-25. Retrieved 26 June 2022.
- ^ Walker TS, Bais HP, Déziel E, Schweizer HP, Rahme LG, Fall R, et al. (January 2004). "Pseudomonas aeruginosa-plant root interactions. Pathogenicity, biofilm formation, and root exudation". Plant Physiology. 134 (1): 320–331. doi:10.1104/pp.103.027888. PMC 316311. PMID 14701912.
- ^ a b Rahme LG, Stevens EJ, Wolfort SF, Shao J, Tompkins RG, Ausubel FM (June 1995). "Common virulence factors for bacterial pathogenicity in plants and animals". Science. 268 (5219): 1899–1902. Bibcode:1995Sci...268.1899R. doi:10.1126/science.7604262. PMID 7604262.
- ^ Rahme LG, Tan MW, Le L, Wong SM, Tompkins RG, Calderwood SB, et al. (November 1997). "Use of model plant hosts to identify Pseudomonas aeruginosa virulence factors". Proceedings of the National Academy of Sciences of the United States of America. 94 (24): 13245–13250. Bibcode:1997PNAS...9413245R. doi:10.1073/pnas.94.24.13245. PMC 24294. PMID 9371831.
- ^ Mahajan-Miklos S, Tan MW, Rahme LG, Ausubel FM (January 1999). "Molecular mechanisms of bacterial virulence elucidated using a Pseudomonas aeruginosa-Caenorhabditis elegans pathogenesis model". Cell. 96 (1): 47–56. doi:10.1016/S0092-8674(00)80958-7. PMID 9989496. S2CID 11207155.
- ^ Martínez C, Pons E, Prats G, León J (January 2004). "Salicylic acid regulates flowering time and links defence responses and reproductive development". The Plant Journal. 37 (2): 209–217. doi:10.1046/j.1365-313X.2003.01954.x. PMID 14690505.
- ^ D'Argenio DA, Gallagher LA, Berg CA, Manoil C (February 2001). "Drosophila as a model host for Pseudomonas aeruginosa infection". Journal of Bacteriology. 183 (4): 1466–1471. doi:10.1128/JB.183.4.1466-1471.2001. PMC 95024. PMID 11157963.
- ^ Miyata S, Casey M, Frank DW, Ausubel FM, Drenkard E (May 2003). "Use of the Galleria mellonella caterpillar as a model host to study the role of the type III secretion system in Pseudomonas aeruginosa pathogenesis". Infection and Immunity. 71 (5): 2404–2413. doi:10.1128/IAI.71.5.2404-2413.2003. PMC 153283. PMID 12704110.
- ^ Rahme LG, Ausubel FM, Cao H, Drenkard E, Goumnerov BC, Lau GW, et al. (August 2000). "Plants and animals share functionally common bacterial virulence factors". Proceedings of the National Academy of Sciences of the United States of America. 97 (16): 8815–8821. Bibcode:2000PNAS...97.8815R. doi:10.1073/pnas.97.16.8815. PMC 34017. PMID 10922040.
- ^ Rumbaugh KP, Griswold JA, Iglewski BH, Hamood AN (November 1999). "Contribution of quorum sensing to the virulence of Pseudomonas aeruginosa in burn wound infections". Infection and Immunity. 67 (11): 5854–5862. doi:10.1128/IAI.67.11.5854-5862.1999. PMC 96966. PMID 10531240.
- ^ Azimi S, Klementiev AD, Whiteley M, Diggle SP (September 2020). "Bacterial Quorum Sensing During Infection". Annual Review of Microbiology. 74 (1). Annual Reviews: 201–219. doi:10.1146/annurev-micro-032020-093845. PMID 32660382. S2CID 220518911.
- ^ Allesen-Holm M, Barken KB, Yang L, Klausen M, Webb JS, Kjelleberg S, et al. (February 2006). "A characterization of DNA release in Pseudomonas aeruginosa cultures and biofilms". Molecular Microbiology. 59 (4): 1114–1128. doi:10.1111/j.1365-2958.2005.05008.x. PMID 16430688. S2CID 11915780.
- ^ a b Dekimpe V, Déziel E (March 2009). "Revisiting the quorum-sensing hierarchy in Pseudomonas aeruginosa: the transcriptional regulator RhlR regulates LasR-specific factors". Microbiology. 155 (Pt 3): 712–723. doi:10.1099/mic.0.022764-0. PMID 19246742.
- ^ Lee J, Zhang L (January 2015). "The hierarchy quorum sensing network in Pseudomonas aeruginosa". Protein & Cell. 6 (1): 26–41. doi:10.1007/s13238-014-0100-x. PMC 4286720. PMID 25249263.
- ^ Winstanley C, Fothergill JL (January 2009). "The role of quorum sensing in chronic cystic fibrosis Pseudomonas aeruginosa infections". FEMS Microbiology Letters. 290 (1): 1–9. doi:10.1111/j.1574-6968.2008.01394.x. PMID 19016870.
- ^ Hoffman LR, Kulasekara HD, Emerson J, Houston LS, Burns JL, Ramsey BW, et al. (January 2009). "Pseudomonas aeruginosa lasR mutants are associated with cystic fibrosis lung disease progression". Journal of Cystic Fibrosis. 8 (1): 66–70. doi:10.1016/j.jcf.2008.09.006. PMC 2631641. PMID 18974024.
- ^ Feltner JB, Wolter DJ, Pope CE, Groleau MC, Smalley NE, Greenberg EP, et al. (October 2016). "LasR Variant Cystic Fibrosis Isolates Reveal an Adaptable Quorum-Sensing Hierarchy in Pseudomonas aeruginosa". mBio. 7 (5): e01513–16, /mbio/7/5/e01513–16.atom. doi:10.1128/mBio.01513-16. PMC 5050340. PMID 27703072.
- ^ a b c Cornelis P (2008). Pseudomonas: Genomics and Molecular Biology (1st ed.). Caister Academic Press. ISBN 978-1-904455-19-6. Archived from the original on 2016-09-12. Retrieved 2007-09-24.
- ^ Bjarnsholt T, Jensen PØ, Rasmussen TB, Christophersen L, Calum H, Hentzer M, et al. (December 2005). "Garlic blocks quorum sensing and promotes rapid clearing of pulmonary Pseudomonas aeruginosa infections". Microbiology. 151 (Pt 12): 3873–3880. doi:10.1099/mic.0.27955-0. PMID 16339933.
- ^ Laventie BJ, Sangermani M, Estermann F, Manfredi P, Planes R, Hug I, et al. (January 2019). "A Surface-Induced Asymmetric Program Promotes Tissue Colonization by Pseudomonas aeruginosa". Cell Host & Microbe. 25 (1): 140–152.e6. doi:10.1016/j.chom.2018.11.008. PMID 30581112.
- ^ Das T, ed. (2021-06-09). Pseudomonas aeruginosa - Biofilm Formation, Infections and Treatments. IntechOpen. doi:10.5772/intechopen.87468. ISBN 978-1-83968-647-4. S2CID 237911377. Archived from the original on 2021-06-22. Retrieved 2024-01-17.
- ^ Thi MT, Wibowo D, Rehm BH (November 2020). "Pseudomonas aeruginosa Biofilms". International Journal of Molecular Sciences. 21 (22): 8671. doi:10.3390/ijms21228671. PMC 7698413. PMID 33212950.
- ^ Ghafoor A, Hay ID, Rehm BH (August 2011). "Role of exopolysaccharides in Pseudomonas aeruginosa biofilm formation and architecture". Applied and Environmental Microbiology. 77 (15): 5238–5246. Bibcode:2011ApEnM..77.5238G. doi:10.1128/AEM.00637-11. PMC 3147449. PMID 21666010.
- ^ Jennings LK, Storek KM, Ledvina HE, Coulon C, Marmont LS, Sadovskaya I, et al. (September 2015). "Pel is a cationic exopolysaccharide that cross-links extracellular DNA in the Pseudomonas aeruginosa biofilm matrix". Proceedings of the National Academy of Sciences of the United States of America. 112 (36): 11353–11358. Bibcode:2015PNAS..11211353J. doi:10.1073/pnas.1503058112. PMC 4568648. PMID 26311845.
- ^ a b Chua SL, Liu Y, Yam JK, Chen Y, Vejborg RM, Tan BG, et al. (July 2014). "Dispersed cells represent a distinct stage in the transition from bacterial biofilm to planktonic lifestyles". Nature Communications. 5: 4462. Bibcode:2014NatCo...5.4462C. doi:10.1038/ncomms5462. PMID 25042103.
- ^ Chua SL, Hultqvist LD, Yuan M, Rybtke M, Nielsen TE, Givskov M, et al. (August 2015). "In vitro and in vivo generation and characterization of Pseudomonas aeruginosa biofilm-dispersed cells via c-di-GMP manipulation". Nature Protocols. 10 (8): 1165–1180. doi:10.1038/nprot.2015.067. hdl:10356/84100. PMID 26158442. S2CID 20235088.
- ^ Ma Y, Deng Y, Hua H, Khoo BL, Chua SL (August 2023). "Distinct bacterial population dynamics and disease dissemination after biofilm dispersal and disassembly". The ISME Journal. 17 (8): 1290–1302. Bibcode:2023ISMEJ..17.1290M. doi:10.1038/s41396-023-01446-5. PMC 10356768. PMID 37270584.
- ^ Mah TF, Pitts B, Pellock B, Walker GC, Stewart PS, O'Toole GA (November 2003). "A genetic basis for Pseudomonas aeruginosa biofilm antibiotic resistance". Nature. 426 (6964): 306–310. Bibcode:2003Natur.426..306M. doi:10.1038/nature02122. PMID 14628055. S2CID 4412747. Archived from the original on 2022-08-17. Retrieved 2022-12-21.
- ^ Wheeler T. "What Is a Pseudomonas Infection?". MedicineNet. Archived from the original on 27 October 2020. Retrieved 8 December 2020.
- ^ "Pseudomonas aeruginosa in Healthcare Settings". Center for Disease Control and Prevention. U.S. Department of Health & Human Services. 6 November 2019. Archived from the original on 29 December 2017. Retrieved 8 December 2020.
- ^ Ryan KJ, Ray CG, eds. (2004). Sherris Medical Microbiology (4th ed.). McGraw Hill. ISBN 978-0-8385-8529-0.
- ^ Iglewski BH (1996). "Pseudomonas". In Baron S, et al. (eds.). Baron's Medical Microbiology (4th ed.). University of Texas Medical Branch. ISBN 978-0-9631172-1-2.
- ^ Anzai Y, Kim H, Park JY, Wakabayashi H, Oyaizu H (July 2000). "Phylogenetic affiliation of the pseudomonads based on 16S rRNA sequence". International Journal of Systematic and Evolutionary Microbiology. 50 (Pt 4): 1563–1589. doi:10.1099/00207713-50-4-1563. PMID 10939664.
- ^ a b c Brenner DJ, Krieg NR, Staley JT, Garrity GM, Boone DR, De Vos P, Goodfellow M, Rainey FA, Schleifer K, eds. (2005). Bergey's manual of systematic bacteriology (2nd ed.). New York: Springer. pp. 323–442. doi:10.1007/0-387-28022-7_9. ISBN 0-387-98771-1. OCLC 45951601. Archived from the original on 2023-03-09. Retrieved 2022-04-21.
- ^ King EO, Ward MK, Raney DE (August 1954). "Two simple media for the demonstration of pyocyanin and fluorescin". The Journal of Laboratory and Clinical Medicine. 44 (2): 301–307. PMID 13184240.
- ^ Striebich RC, Smart CE, Gunasekera TS, Mueller SS, Strobel EM, McNichols BW, et al. (September 2014). "Characterization of the F-76 diesel and Jet-A aviation fuel hydrocarbon degradation profiles of Pseudomonas aeruginosa and Marinobacter hydrocarbonoclasticus". International Biodeterioration & Biodegradation. 93: 33–43. Bibcode:2014IBiBi..93...33S. doi:10.1016/j.ibiod.2014.04.024.
- ^ Hachem RY, Chemaly RF, Ahmar CA, Jiang Y, Boktour MR, Rjaili GA, et al. (June 2007). "Colistin is effective in treatment of infections caused by multidrug-resistant Pseudomonas aeruginosa in cancer patients". Antimicrobial Agents and Chemotherapy. 51 (6): 1905–1911. doi:10.1128/AAC.01015-06. PMC 1891378. PMID 17387153.
- ^ Nagoba BS, Selkar SP, Wadher BJ, Gandhi RC (December 2013). "Acetic acid treatment of pseudomonal wound infections--a review". Journal of Infection and Public Health. 6 (6): 410–415. doi:10.1016/j.jiph.2013.05.005. PMID 23999348.
- ^ Barceló IM, Torrens G, Escobar-Salom M, Jordana-Lluch E, Capó-Bauzá MM, Ramón-Pallín C, et al. (February 2022). "Impact of Peptidoglycan Recycling Blockade and Expression of Horizontally Acquired β-Lactamases on Pseudomonas aeruginosa Virulence". Microbiology Spectrum. 10 (1): e0201921. doi:10.1128/spectrum.02019-21. PMC 8849096. PMID 35171032.
- ^ Mirsalehian A, Kalantar-Neyestanaki D, Nourijelyani K, Asadollahi K, Taherikalani M, Emaneini M, et al. (October 2014). "Detection of AmpC-β-lactamases producing isolates among carbapenem resistant P. aeruginosa isolated from burn patient". Iranian Journal of Microbiology. 6 (5): 306–310. PMC 4385569. PMID 25848519.
- ^ Atilla A, Eroğlu C, Esen S, Sünbül M, Leblebicioğlu H (January 2012). "[Investigation of the frequency of PER-1 type beta-lactamase and antimicrobial resistance rates in nosocomial isolates of Pseudomonas aeruginosa]". Mikrobiyoloji Bulteni (in Turkish). 46 (1): 1–8. PMID 22399165.
- ^ Evans BA, Amyes SG (April 2014). "OXA β-lactamases". Clinical Microbiology Reviews. 27 (2): 241–263. doi:10.1128/CMR.00117-13. PMC 3993105. PMID 24696435.
- ^ Shaikh S, Fatima J, Shakil S, Rizvi SM, Kamal MA (January 2015). "Antibiotic resistance and extended spectrum beta-lactamases: Types, epidemiology and treatment". Saudi Journal of Biological Sciences. 22 (1): 90–101. doi:10.1016/j.sjbs.2014.08.002. PMC 4281622. PMID 25561890.
- ^ Hirakata Y, Yamaguchi T, Nakano M, Izumikawa K, Mine M, Aoki S, et al. (July 2003). "Clinical and bacteriological characteristics of IMP-type metallo-beta-lactamase-producing Pseudomonas aeruginosa". Clin Infect Dis. 37 (1): 26–32. doi:10.1086/375594. PMID 12830405.
- ^ Pagani L, Colinon C, Migliavacca R, Labonia M, Docquier JD, Nucleo E, et al. (August 2005). "Nosocomial outbreak caused by multidrug-resistant Pseudomonas aeruginosa producing IMP-13 metallo-beta-lactamase". J Clin Microbiol. 43 (8): 3824–8. doi:10.1128/JCM.43.8.3824-3828.2005. PMC 1233900. PMID 16081918.
- ^ Kiyaga S, Kyany'a C, Muraya AW, Smith HJ, Mills EG, Kibet C, et al. (2022). "Genetic Diversity, Distribution, and Genomic Characterization of Antibiotic Resistance and Virulence of Clinical Pseudomonas aeruginosa Strains in Kenya". Frontiers in Microbiology. 13: 835403. doi:10.3389/fmicb.2022.835403. PMC 8964364. PMID 35369511.
- ^ Khan AU, Maryam L, Zarrilli R (April 2017). "Structure, Genetics and Worldwide Spread of New Delhi Metallo-β-lactamase (NDM): a threat to public health". BMC Microbiology. 17 (1): 101. doi:10.1186/s12866-017-1012-8. PMC 5408368. PMID 28449650.
- ^ Dortet L, Poirel L, Nordmann P (2014). "Worldwide dissemination of the NDM-type carbapenemases in Gram-negative bacteria". BioMed Research International. 2014: 249856. doi:10.1155/2014/249856. PMC 3984790. PMID 24790993.
- ^ Docquier JD, Lamotte-Brasseur J, Galleni M, Amicosante G, Frère JM, Rossolini GM (February 2003). "On functional and structural heterogeneity of VIM-type metallo-beta-lactamases". The Journal of Antimicrobial Chemotherapy. 51 (2): 257–266. doi:10.1093/jac/dkg067. PMID 12562689.
- ^ Galimand M, Lambert T, Gerbaud G, Courvalin P (July 1993). "Characterization of the aac(6')-Ib gene encoding an aminoglycoside 6'-N-acetyltransferase in Pseudomonas aeruginosa BM2656". Antimicrobial Agents and Chemotherapy. 37 (7): 1456–1462. doi:10.1128/AAC.37.7.1456. PMC 187994. PMID 8363376.
- ^ Coban AY, Tanrıverdi Çaycı Y, Yıldırım T, Erturan Z, Durupınar B, Bozdoğan B (October 2011). "[Investigation of plasmid-mediated quinolone resistance in Pseudomonas aeruginosa strains isolated from cystic fibrosis patients]". Mikrobiyoloji Bulteni (in Turkish). 45 (4): 602–608. PMID 22090290.
- ^ Mielko KA, Jabłoński SJ, Milczewska J, Sands D, Łukaszewicz M, Młynarz P (November 2019). "Metabolomic studies of Pseudomonas aeruginosa". World Journal of Microbiology & Biotechnology. 35 (11): 178. doi:10.1007/s11274-019-2739-1. PMC 6838043. PMID 31701321.
- ^ Jurado-Martín I, Sainz-Mejías M, McClean S (March 2021). "Pseudomonas aeruginosa: An Audacious Pathogen with an Adaptable Arsenal of Virulence Factors". International Journal of Molecular Sciences. 22 (6): 3128. doi:10.3390/ijms22063128. PMC 8003266. PMID 33803907.
- ^ Paul SI, Rahman A, Rahman MM (January 2023). Baltrus DA (ed.). "Friend or Foe: Whole-Genome Sequence of Pseudomonas aeruginosa TG523, Isolated from the Gut of a Healthy Nile Tilapia (Oreochromis niloticus)". Microbiology Resource Announcements. 12 (1): e0113322. doi:10.1128/mra.01133-22. PMC 9872575. PMID 36598220.
- ^ Poole K (January 2004). "Efflux-mediated multiresistance in Gram-negative bacteria". Clinical Microbiology and Infection. 10 (1): 12–26. doi:10.1111/j.1469-0691.2004.00763.x. PMID 14706082.
- ^ Rampioni G, Pillai CR, Longo F, Bondì R, Baldelli V, Messina M, et al. (September 2017). "Effect of efflux pump inhibition on Pseudomonas aeruginosa transcriptome and virulence". Scientific Reports. 7 (1): 11392. Bibcode:2017NatSR...711392R. doi:10.1038/s41598-017-11892-9. PMC 5596013. PMID 28900249.
- ^ Aghapour Z, Gholizadeh P, Ganbarov K, Bialvaei AZ, Mahmood SS, Tanomand A, et al. (2019). "Molecular mechanisms related to colistin resistance in Enterobacteriaceae". Infection and Drug Resistance. 12: 965–975. doi:10.2147/IDR.S199844. PMC 6519339. PMID 31190901.
- ^ Wong A, Rodrigue N, Kassen R (September 2012). "Genomics of adaptation during experimental evolution of the opportunistic pathogen Pseudomonas aeruginosa". PLOS Genetics. 8 (9): e1002928. doi:10.1371/journal.pgen.1002928. PMC 3441735. PMID 23028345.
- ^ a b Zhang YF, Han K, Chandler CE, Tjaden B, Ernst RK, Lory S (December 2017). "Probing the sRNA regulatory landscape of P. aeruginosa: post-transcriptional control of determinants of pathogenicity and antibiotic susceptibility". Molecular Microbiology. 106 (6): 919–937. doi:10.1111/mmi.13857. PMC 5738928. PMID 28976035.
- ^ Kawasaki K, China K, Nishijima M (July 2007). "Release of the lipopolysaccharide deacylase PagL from latency compensates for a lack of lipopolysaccharide aminoarabinose modification-dependent resistance to the antimicrobial peptide polymyxin B in Salmonella enterica". Journal of Bacteriology. 189 (13): 4911–4919. doi:10.1128/JB.00451-07. PMC 1913436. PMID 17483225.
- ^ Forestier C, Guelon D, Cluytens V, Gillart T, Sirot J, De Champs C (2008). "Oral probiotic and prevention of Pseudomonas aeruginosa infections: a randomized, double-blind, placebo-controlled pilot study in intensive care unit patients". Critical Care. 12 (3): R69. doi:10.1186/cc6907. PMC 2481460. PMID 18489775.[non-primary source needed]
- ^ Döring G, Pier GB (February 2008). "Vaccines and immunotherapy against Pseudomonas aeruginosa". Vaccine. 26 (8): 1011–1024. doi:10.1016/j.vaccine.2007.12.007. PMID 18242792.
- ^ "Pseudomonas Aeruginosa Fact Sheet" (PDF). Children's Hospital of Illinois. Archived from the original (PDF) on 2016-05-09. Retrieved 2014-11-15.
- ^ Sulakvelidze A, Alavidze Z, Morris JG (March 2001). "Bacteriophage therapy". Antimicrobial Agents and Chemotherapy. 45 (3): 649–659. doi:10.1128/AAC.45.3.649-659.2001. PMC 90351. PMID 11181338.
- ^ Wright A, Hawkins CH, Anggård EE, Harper DR (August 2009). "A controlled clinical trial of a therapeutic bacteriophage preparation in chronic otitis due to antibiotic-resistant Pseudomonas aeruginosa; a preliminary report of efficacy". Clinical Otolaryngology. 34 (4): 349–357. doi:10.1111/j.1749-4486.2009.01973.x. PMID 19673983. S2CID 379471.
- ^ Ipoutcha T, Racharaks R, Huttelmaier S, Wilson CJ, Ozer EA, Hartmann EM (31 January 2024). "A synthetic biology approach to assemble and reboot clinically relevant Pseudomonas aeruginosa tailed phages". Microbiology Spectrum. 12 (3): e0289723. doi:10.1128/spectrum.02897-23. PMC 10913387. PMID 38294230.
- ^ van Ditmarsch D, Boyle KE, Sakhtah H, Oyler JE, Nadell CD, Déziel É, et al. (August 2013). "Convergent evolution of hyperswarming leads to impaired biofilm formation in pathogenic bacteria". Cell Reports. 4 (4): 697–708. doi:10.1016/j.celrep.2013.07.026. PMC 3770465. PMID 23954787.
- ^ Zimmer C (15 August 2013). "Watching Bacteria Evolve, With Predictable Results". The New York Times. Archived from the original on 2 January 2018. Retrieved 2 February 2016.
- ^ Pathak VM (23 March 2017). "Review on the current status of polymer degradation: a microbial approach". Bioresources and Bioprocessing. 4: 15. doi:10.1186/s40643-017-0145-9. ISSN 2197-4365.
- ^ Payne DD, Renz A, Dunphy LJ, Lewis T, Dräger A, Papin JA (October 2021). "An updated genome-scale metabolic network reconstruction of Pseudomonas aeruginosa PA14 to characterize mucin-driven shifts in bacterial metabolism". npj Systems Biology and Applications. 7 (1): 37. doi:10.1038/s41540-021-00198-2. PMC 8501023. PMID 34625561. S2CID 232224404.
- ^ Dahal S, Renz A, Dräger A, Yang L (February 2023). "Genome-scale model of Pseudomonas aeruginosa metabolism unveils virulence and drug potentiation". Communications Biology. 6 (1): 165. doi:10.1038/s42003-023-04540-8. PMC 9918512. PMID 36765199. S2CID 256702200.
- ^ East African Community (2019-11-29). Pest Risk Analysis (PRA) for Grain and Seed of Beans, Phaseolus vulgaris L. within East African Countries (Kenya, Burundi, Rwanda, Tanzania and Uganda): A Qualitative, Pathway-Initiated Risk Analysis) (Report). hdl:11671/24138. Archived from the original on 2022-05-26. Retrieved 2021-10-19.
- ^ "Infection toll for recalled eyedrops climbs to 81, including 4 deaths, CDC says". NPR. Archived from the original on 2023-05-29.
Further reading
[edit]- Sweere JM, Van Belleghem JD, Ishak H, Bach MS, Popescu M, Sunkari V, et al. (March 2019). "Bacteriophage trigger antiviral immunity and prevent clearance of bacterial infection". Science. 363 (6434). doi:10.1126/science.aat9691. PMC 6656896. PMID 30923196. about Pseudomonas aeruginosa filamentous phages (Pf-phages), Inoviridae. See also:
External links
[edit]- "Pseudomonas aeruginosa DSM 50071". The Bacterial Diversity Metadatabase.
