Molecular genetics
This article needs additional citations for verification. (January 2012) |
| Part of a series on |
| Genetics |
|---|
 |
Molecular genetics is a branch of biology that addresses how differences in the structures or expression of DNA molecules manifests as variation among organisms. Molecular genetics often applies an "investigative approach" to determine the structure and/or function of genes in an organism's genome using genetic screens.[1][2]
The field of study is based on the merging of several sub-fields in biology: classical Mendelian inheritance, cellular biology, molecular biology, biochemistry, and biotechnology. It integrates these disciplines to explore things like genetic inheritance, gene regulation and expression, and the molecular mechanism behind various life processes.[1]
A key goal of molecular genetics is to identify and study genetic mutations. Researchers search for mutations in a gene or induce mutations in a gene to link a gene sequence to a specific phenotype.[3] Therefore molecular genetics is a powerful methodology for linking mutations to genetic conditions that may aid the search for treatments of various genetics diseases.
History
[edit]The discovery of DNA as the blueprint for life and breakthroughs in molecular genetics research came from the combined works of many scientists. In 1869, chemist Johann Friedrich Miescher, who was researching the composition of white blood cells, discovered and isolated a new molecule that he named nuclein from the cell nucleus, which would ultimately be the first discovery of the molecule DNA that was later determined to be the molecular basis of life. He determined it was composed of hydrogen, oxygen, nitrogen and phosphorus.[4] Biochemist Albrecht Kossel identified nuclein as a nucleic acid and provided its name deoxyribonucleic acid (DNA). He continued to build on that by isolating the basic building blocks of DNA and RNA; made up of the nucleotides: adenine, guanine, thymine, cytosine. and uracil. His work on nucleotides earned him a Nobel Prize in Physiology.[5]
In the early 1900s, Gregor Mendel, who became known as one of the fathers of genetics, made great contributions to the field of genetics through his various experiments with pea plants where he was able to discover the principles of inheritance such as recessive and dominant traits, without knowing what genes where composed of.[6] In the mid 19th century, anatomist Walther Flemming, discovered what we now know as chromosomes and the separation process they undergo through mitosis. His work along with Theodor Boveri first came up with the Chromosomal Theory of Inheritance, which helped explain some of the patterns Mendel had observed much earlier.[7]
For molecular genetics to develop as a discipline, several scientific discoveries were necessary. The discovery of DNA as a means to transfer the genetic code of life from one cell to another and between generations was essential for identifying the molecule responsible for heredity. Molecular genetics arose initially from studies involving genetic transformation in bacteria. In 1944 Avery, McLeod and McCarthy[8] isolated DNA from a virulent strain of S. pneumoniae, and using just this DNA were able to convert a harmless strain to virulence. They called the uptake, incorporation and expression of DNA by bacteria "transformation". This finding suggested that DNA is the genetic material of bacteria. [9] Bacterial transformation is often induced by conditions of stress, and the function of transformation appears to be repair of genomic damage.[9]
In 1950, Erwin Chargaff derived rules that offered evidence of DNA being the genetic material of life. These were "1) that the base composition of DNA varies between species and 2) in natural DNA molecules, the amount of adenine (A) is equal to the amount of thymine (T), and the amount of guanine (G) is equal to the amount of cytosine (C)."[10] These rules, known as Chargaff's rules, helped to understand of molecular genetics.[10] In 1953 Francis Crick and James Watson, building upon the X-ray crystallography work done by Rosalind Franklin and Maurice Wilkins, were able to derive the 3-D double helix structure of DNA.[11]
The phage group was an informal network of biologists centered on Max Delbrück that contributed substantially to molecular genetics and the origins of molecular biology during the period from about 1945 to 1970.[12] The phage group took its name from bacteriophages, the bacteria-infecting viruses that the group used as experimental model organisms. Studies by molecular geneticists affiliated with this group contributed to understanding how gene-encoded proteins function in DNA replication, DNA repair and DNA recombination, and on how viruses are assembled from protein and nucleic acid components (molecular morphogenesis). Furthermore, the role of chain terminating codons was elucidated. One noteworthy study was performed by Sydney Brenner and collaborators using "amber" mutants defective in the gene encoding the major head protein of bacteriophage T4.[13] This study demonstrated the co-linearity of the gene with its encoded polypeptide, thus providing strong evidence for the "sequence hypothesis" that the amino acid sequence of a protein is specified by the nucleotide sequence of the gene determining the protein.
The isolation of a restriction endonuclease in E. coli by Arber and Linn in 1969 opened the field of genetic engineering.[14] Restriction enzymes were used to linearize DNA for separation by electrophoresis and Southern blotting allowed for the identification of specific DNA segments via hybridization probes.[15][16] In 1971, Berg utilized restriction enzymes to create the first recombinant DNA molecule and first recombinant DNA plasmid.[17] In 1972, Cohen and Boyer created the first recombinant DNA organism by inserting recombinant DNA plasmids into E. coli, now known as bacterial transformation, and paved the way for molecular cloning.[18] The development of DNA sequencing techniques in the late 1970s, first by Maxam and Gilbert, and then by Frederick Sanger, was pivotal to molecular genetic research and enabled scientists to begin conducting genetic screens to relate genotypic sequences to phenotypes.[19] Polymerase chain reaction (PCR) using Taq polymerase, invented by Mullis in 1985, enabled scientists to create millions of copies of a specific DNA sequence that could be used for transformation or manipulated using agarose gel separation.[20] A decade later, the first whole genome was sequenced (Haemophilus influenzae), followed by the eventual sequencing of the human genome via the Human Genome Project in 2001.[21] The culmination of all of those discoveries was a new field called genomics that links the molecular structure of a gene to the protein or RNA encoded by that segment of DNA and the functional expression of that protein within an organism.[22] Today, through the application of molecular genetic techniques, genomics is being studied in many model organisms and data is being collected in computer databases like NCBI and Ensembl. The computer analysis and comparison of genes within and between different species is called bioinformatics, and links genetic mutations on an evolutionary scale.[23]
Central dogma
[edit]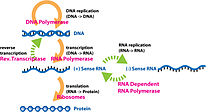
The central dogma plays a key role in the study of molecular genetics. The central dogma states that DNA replicates itself, DNA is transcribed into RNA, and RNA is translated into proteins.[24] Along with the central dogma, the genetic code is used in understanding how RNA is translated into proteins. Replication of DNA and transcription from DNA to mRNA occurs in the nucleus while translation from RNA to proteins occurs in the ribosome.[25] The genetic code is made of four interchangeable parts othe DNA molecules, called "bases": adenine, cytosine, uracil (in RNA; thymine in DNA), and guanine and is redundant, meaning multiple combinations of these base pairs (which are read in triplicate) produce the same amino acid.[26] Proteomics and genomics are fields in biology that come out of the study of molecular genetics and the central dogma.[27]
Structure of DNA
[edit]An organism's genome is made up by its entire set of DNA and is responsible for its genetic traits, function and development. The composition of DNA itself is an essential component to the field of molecular genetics; it is the basis of how DNA is able to store genetic information, pass it on, and be in a format that can be read and translated.[28]
DNA is a double stranded molecule, with each strand oriented in an antiparallel fashion. Nucleotides are the building blocks of DNA, each composed of a sugar molecule, a phosphate group and one of four nitrogenous bases: adenine, guanine, cytosine, and thymine. A single strand of DNA is held together by covalent bonds, while the two antiparallel strands are held together by hydrogen bonds between the nucleotide bases. Adenine binds with thymine and cytosine binds with guanine. It is these four base sequences that form the genetic code for all biological life and contains the information for all the proteins the organism will be able to synthesize.[29]
Its unique structure allows DNA to store and pass on biological information across generations during cell division. At cell division, cells must be able to copy its genome and pass it on to daughter cells. This is possible due to the double-stranded structure of DNA because one strand is complementary to its partner strand, and therefore each of these strands can act as a template strand for the formation of a new complementary strand. This is why the process of DNA replication is known as a semiconservative process.[30]
Techniques
[edit]Forward genetics
[edit]Forward genetics is a molecular genetics technique used to identify genes or genetic mutations that produce a certain phenotype. In a genetic screen, random mutations are generated with mutagens (chemicals or radiation) or transposons and individuals are screened for the specific phenotype. Often, a secondary assay in the form of a selection may follow mutagenesis where the desired phenotype is difficult to observe, for example in bacteria or cell cultures. The cells may be transformed using a gene for antibiotic resistance or a fluorescent reporter so that the mutants with the desired phenotype are selected from the non-mutants.[31]
Mutants exhibiting the phenotype of interest are isolated and a complementation test may be performed to determine if the phenotype results from more than one gene. The mutant genes are then characterized as dominant (resulting in a gain of function), recessive (showing a loss of function), or epistatic (the mutant gene masks the phenotype of another gene). Finally, the location and specific nature of the mutation is mapped via sequencing.[32] Forward genetics is an unbiased approach and often leads to many unanticipated discoveries, but may be costly and time consuming. Model organisms like the nematode worm Caenorhabditis elegans, the fruit fly Drosophila melanogaster, and the zebrafish Danio rerio have been used successfully to study phenotypes resulting from gene mutations.[33]

Reverse genetics
[edit]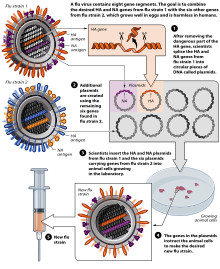
Reverse genetics is the term for molecular genetics techniques used to determine the phenotype resulting from an intentional mutation in a gene of interest. The phenotype is used to deduce the function of the un-mutated version of the gene. Mutations may be random or intentional changes to the gene of interest. Mutations may be a missense mutation caused by nucleotide substitution, a nucleotide addition or deletion to induce a frameshift mutation, or a complete addition/deletion of a gene or gene segment. The deletion of a particular gene creates a gene knockout where the gene is not expressed and a loss of function results (e.g. knockout mice). Missense mutations may cause total loss of function or result in partial loss of function, known as a knockdown. Knockdown may also be achieved by RNA interference (RNAi).[35] Alternatively, genes may be substituted into an organism's genome (also known as a transgene) to create a gene knock-in and result in a gain of function by the host.[36] Although these techniques have some inherent bias regarding the decision to link a phenotype to a particular function, it is much faster in terms of production than forward genetics because the gene of interest is already known.
Molecular genetic tools
[edit]Molecular genetics is a scientific approach that utilizes the fundamentals of genetics as a tool to better understand the molecular basis of a disease and biological processes in organisms. Below are some tools readily employed by researchers in the field.
Microsatellites
[edit]Microsatellites or single sequence repeats (SSRS) are short repeating segment of DNA composed to 6 nucleotides at a particular location on the genome that are used as genetic marker. Researchers can analyze these microsatellites in techniques such DNA fingerprinting and paternity testing since these repeats are highly unique to individuals/families. a can also be used in constructing genetic maps and to studying genetic linkage to locate the gene or mutation responsible for specific trait or disease. Microsatellites can also be applied to population genetics to study comparisons between groups.[37]
Genome-wide association studies
[edit]Genome-wide association studies (GWAS) are a technique that relies on single nucleotide polymorphisms (SNPs) to study genetic variations in populations that can be associated with a particular disease. The Human Genome Project mapped the entire human genome and has made this approach more readily available and cost effective for researchers to implement. In order to conduct a GWAS researchers use two groups, one group that has the disease researchers are studying and another that acts as the control that does not have that particular disease. DNA samples are obtained from participants and their genome can then be derived through lab machinery and quickly surveyed to compare participants and look for SNPs that can potentially be associated with the disease. This technique allows researchers to pinpoint genes and locations of interest in the human genome that they can then further study to identify that cause of the disease.[38]
Karyotyping
[edit]Karyotyping allows researchers to analyze chromosomes during metaphase of mitosis, when they are in a condensed state. Chromosomes are stained and visualized through a microscope to look for any chromosomal abnormalities. This technique can be used to detect congenital genetic disorder such as down syndrome, identify gender in embryos, and diagnose some cancers that are caused by chromosome mutations such as translocations.[39]
Modern applications
[edit]Genetic engineering
[edit]Genetic engineering is an emerging field of science, and researcher are able to leverage molecular genetic technology to modify the DNA of organisms and create genetically modified and enhanced organisms for industrial, agricultural and medical purposes. This can be done through genome editing techniques, which can involve modifying base pairings in a DNA sequence, or adding and deleting certain regions of DNA.[40]
Gene editing
[edit]Gene editing allows scientists to alter/edit an organism's DNA. One way to due this is through the technique Crispr/Cas9, which was adapted from the genome immune defense that is naturally occurring in bacteria. This technique relies on the protein Cas9 which allows scientists to make a cut in strands of DNA at a specific location, and it uses a specialized RNA guide sequence to ensure the cut is made in the proper location in the genome. Then scientists use DNAs repair pathways to induce changes in the genome; this technique has wide implications for disease treatment.[41]
Personalized medicine
[edit]Molecular genetics has wide implications in medical advancement and understanding the molecular basis of a disease allows the opportunity for more effective diagnostic and therapies. One of the goals of the field is personalized medicine, where an individual's genetics can help determine the cause and tailor the cure for a disease they are afflicted with and potentially allow for more individualized treatment approaches which could be more effective. For example, certain genetic variations in individuals could make them more receptive to a particular drug while other could have a higher risk of adverse reaction to treatments. So this information would allow researchers and clinicals to make the most informed decisions about treatment efficacy for patients rather than the standard trial and error approach.[42]
Forensic genetics
[edit]Forensic genetics plays an essential role for criminal investigations through that use of various molecular genetic techniques. One common technique is DNA fingerprinting which is done using a combination of molecular genetic techniques like polymerase chain reaction (PCR) and gel electrophoresis. PCR is a technique that allows a target DNA sequence to be amplified, meaning even a tiny quantity of DNA from a crime scene can be extracted and replicated many times to provide a sufficient amount of material for analysis. Gel electrophoresis allows the DNA sequence to be separated based on size, and the pattern that is derived is known as DNA fingerprinting and is unique to each individual. This combination of molecular genetic techniques allows a simple DNA sequence to be extracted, amplified, analyzed and compared with others and is a standard technique used in forensics.[43]
See also
[edit]- Complementation (genetics)
- DNA damage (naturally occurring)
- DNA damage theory of aging
- Epigenetics
- Gene mapping
- Genetic code
- Genetic recombination
- Genomic imprinting
- History of genetics
- Homologous recombination
- Mutagenesis
- Regulation of gene expression
- Timeline of the history of genetics
- Transformation (genetics)
Sources and notes
[edit]- ^ a b Waters, Ken (2013), "Molecular Genetics", in Zalta, Edward N. (ed.), The Stanford Encyclopedia of Philosophy (Fall 2013 ed.), Metaphysics Research Lab, Stanford University, retrieved 2019-10-07
- ^ Alberts, Bruce (2014-11-18). Molecular biology of the cell (Sixth ed.). New York, NY. ISBN 978-0-8153-4432-2. OCLC 887605755.
{{cite book}}: CS1 maint: location missing publisher (link) - ^ Brown, Terence A. (2002), "Mutation, Repair and Recombination", Genomes. 2nd edition, Wiley-Liss, retrieved 2023-10-09
- ^ Lamm, Ehud; Harman, Oren; Veigl, Sophie Juliane (June 2020). "Before Watson and Crick in 1953 Came Friedrich Miescher in 1869". Genetics. 215 (2): 291–296. doi:10.1534/genetics.120.303195. ISSN 0016-6731. PMC 7268995. PMID 32487691.
- ^ "The Nobel Prize in Physiology or Medicine 1910". NobelPrize.org. Retrieved 2023-10-15.
- ^ "Gregor Mendel and the Principles of Inheritance | Learn Science at Scitable". www.nature.com. Retrieved 2023-10-15.
- ^ Paweletz, N. (January 2001). "Walther Flemming: pioneer of mitosis research". Nature Reviews. Molecular Cell Biology. 2 (1): 72–75. doi:10.1038/35048077. ISSN 1471-0072. PMID 11413469. S2CID 205011982.
- ^ Avery OT, Macleod CM, McCarty M. Studies on the chemical nature of the substance inducing transformation of pneumococcal types: Induction of transformation by a desoxyribonucleic acid fraction isolated form pneumococcus type III. J Exp Med. 1944 Feb 1;79(2):137-58. doi:10.1084/jem.79.2.137. PMID 19871359; PMC 2135445
- ^ a b Bernstein H, Bernstein C, Michod RE (2018). Sex in microbial pathogens. Infection, Genetics and Evolution volume 57, pages 8-25. doi:10.1016/j.meegid.2017.10.024
- ^ a b Elson, David; Chargaff, Erwin (May 1954). "Regularities in the Composition of Pentose Nucleic Acids". Nature. 173 (4413): 1037–1038. Bibcode:1954Natur.173.1037E. doi:10.1038/1731037a0. ISSN 1476-4687. PMID 13165710. S2CID 4219775.
- ^ Watson, J. D.; Crick, F. H. C. (April 1953). "Molecular Structure of Nucleic Acids: A Structure for Deoxyribose Nucleic Acid". Nature. 171 (4356): 737–738. Bibcode:1953Natur.171..737W. doi:10.1038/171737a0. ISSN 1476-4687. PMID 13054692. S2CID 4253007.
- ^ Phage and the Origins of Molecular Biology (2007) Edited by John Cairns, Gunther S. Stent, and James D. Watson, Cold Spring Harbor Laboratory of Quantitative Biology, Cold Spring Harbor, Long Island, New York ISBN 978-0-87969-800-3
- ^ Sarabhai AS, Stretton AO, Brenner S, Bolle A (January 1964). "Co-linearity of the gene with the polypeptide chain". Nature. 201 (4914): 13–7. Bibcode:1964Natur.201...13S. doi:10.1038/201013a0. PMID 14085558. S2CID 10179456
- ^ "Restriction Enzymes Spotlight | Learn Science at Scitable". www.nature.com. Retrieved 2019-10-07.
- ^ Righetti, Pier Giorgio (June 24, 2005). "Electrophoresis: The march of pennies, the march of dimes". Journal of Chromatography A. 1079 (1–2): 24–40. doi:10.1016/j.chroma.2005.01.018. PMID 16038288.
- ^ "Southern Blotting | MyBioSource Learning Center". Retrieved 2019-11-11.
- ^ "Professor Paul Berg | Biographical summary". WhatisBiotechnology.org. Retrieved 2019-10-07.
- ^ "Herbert W. Boyer and Stanley N. Cohen". Science History Institute. 2016-06-01. Retrieved 2019-10-07.
- ^ "DNA sequencing | genetics". Encyclopedia Britannica. Retrieved 2019-10-07.
- ^ "The Invention of PCR". Bitesize Bio. 2007-10-24. Retrieved 2019-10-07.
- ^ "Timeline: Organisms that have had their genomes sequenced". yourgenome. Retrieved 2019-10-07.
- ^ "What is genomics?". EMBL-EBI Train online. 2011-09-09. Retrieved 2019-10-07.
- ^ "What is bioinformatics? A proposed definition and overview of the field". Methods of Information in Medicine. 40 (2). 2001. doi:10.1055/s-008-38405. ISSN 0026-1270.
- ^ "The Central Dogma | Protocol". www.jove.com. Retrieved 2020-12-04.
- ^ "Transcription, Translation and Replication". www.atdbio.com. Retrieved 2020-12-04.
- ^ "Genetic Code". Genome.gov. Retrieved 2020-12-04.
- ^ "A Brief Guide to Genomics". Genome.gov. Retrieved 2020-12-04.
- ^ "Home - Genome - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2023-10-16.
- ^ Alberts, Bruce; Johnson, Alexander; Lewis, Julian; Raff, Martin; Roberts, Keith; Walter, Peter (2002), "The Structure and Function of DNA", Molecular Biology of the Cell. 4th edition, Garland Science, retrieved 2023-10-16
- ^ "Semi-Conservative DNA Replication | Learn Science at Scitable". www.nature.com. Retrieved 2023-10-16.
- ^ "Selection versus Screening in Directed Evolution", Directed Evolution of Selective Enzymes, John Wiley & Sons, Ltd, 2016, pp. 27–57, doi:10.1002/9783527655465.ch2, ISBN 978-3-527-65546-5
- ^ Schneeberger, Korbinian (August 20, 2014). "Using next-generation sequencing to isolate mutant genes from forward genetic screens". Nature Reviews Genetics. 15 (10): 662–676. doi:10.1038/nrg3745. hdl:11858/00-001M-0000-0024-CF80-4. ISSN 1471-0056. PMID 25139187. S2CID 1822657.
- ^ Lawson, Nathan D.; Wolfe, Scot A. (2011-07-19). "Forward and Reverse Genetic Approaches for the Analysis of Vertebrate Development in the Zebrafish". Developmental Cell. 21 (1): 48–64. doi:10.1016/j.devcel.2011.06.007. ISSN 1534-5807. PMID 21763608.
- ^ Kutscher, Lena M. (2014). "Forward and reverse mutagenesis in C. elegans". WormBook: 1–26. doi:10.1895/wormbook.1.167.1. PMC 4078664. PMID 24449699.
- ^ Hardy, Serge; Legagneux, Vincent; Audic, Yann; Paillard, Luc (October 2010). "Reverse genetics in eukaryotes". Biology of the Cell. 102 (10): 561–580. doi:10.1042/BC20100038. PMC 3017359. PMID 20812916.
- ^ Doyle, Alfred; McGarry, Michael P.; Lee, Nancy A.; Lee, James J. (April 2012). "The construction of transgenic and gene knockout/knockin mouse models of human disease". Transgenic Research. 21 (2): 327–349. doi:10.1007/s11248-011-9537-3. ISSN 0962-8819. PMC 3516403. PMID 21800101.
- ^ "Microsatellite". Genome.gov. Retrieved 2023-12-07.
- ^ "Genome-Wide Association Studies Fact Sheet". Genome.gov. Retrieved 2023-12-07.
- ^ "Karyotyping | Learn Science at Scitable". www.nature.com. Retrieved 2023-12-07.
- ^ Howard, Heidi C.; van El, Carla G.; Forzano, Francesca; Radojkovic, Dragica; Rial-Sebbag, Emmanuelle; de Wert, Guido; Borry, Pascal; Cornel, Martina C. (January 2018). "One small edit for humans, one giant edit for humankind? Points and questions to consider for a responsible way forward for gene editing in humans". European Journal of Human Genetics. 26 (1): 1–11. doi:10.1038/s41431-017-0024-z. ISSN 1018-4813. PMC 5839051. PMID 29192152.
- ^ "What are genome editing and CRISPR-Cas9?: MedlinePlus Genetics". medlineplus.gov. Retrieved 2023-12-09.
- ^ Goetz, Laura H.; Schork, Nicholas J. (June 2018). "Personalized Medicine: Motivation, Challenges and Progress". Fertility and Sterility. 109 (6): 952–963. doi:10.1016/j.fertnstert.2018.05.006. ISSN 0015-0282. PMC 6366451. PMID 29935653.
- ^ Li, Chengtao (2018-07-18). "Forensic genetics". Forensic Sciences Research. 3 (2): 103–104. doi:10.1080/20961790.2018.1489445. ISSN 2096-1790. PMC 6197140. PMID 30483657.
Further reading
[edit]- Sites and databases related to genetics, cytogenetics and oncology, at Atlas of Genetics and Cytogenetics in Oncology and Haematology
- Jeremy W. Dale and Simon F. Park. 2010. Molecular Genetics of Bacteria, 5th Edition ISBN 978-0-470-74184-9
External links
[edit] Media related to Molecular genetics at Wikimedia Commons
Media related to Molecular genetics at Wikimedia Commons
