Split gene theory
This article needs attention from an expert in Biology. The specific problem is: Highly technical subject whose notability and content need to be assessed. (January 2019) |
The split gene theory is a theory of the origin of introns, long non-coding sequences in eukaryotic genes between the exons.[1][2][3] The theory holds that the randomness of primordial DNA sequences would only permit small (< 600bp) open reading frames (ORFs), and that important intron structures and regulatory sequences are derived from stop codons. In this introns-first framework, the spliceosomal machinery and the nucleus evolved due to the necessity to join these ORFs (now "exons") into larger proteins, and that intronless bacterial genes are less ancestral than the split eukaryotic genes. The theory originated with Periannan Senapathy.
The theory provides solutions to key questions concerning the split gene architecture, including split eukaryotic genes, exons, introns, splice junctions, and branch points, based on the origin of split genes from random genetic sequences. It also provides possible solutions to the origin of the spliceosomal machinery, the nuclear boundary and the eukaryotic cell.
This theory led to the Shapiro–Senapathy algorithm, which provides the methodology for detecting the splice sites, exons and split genes in eukaryotic DNA, and which is the main method for detecting splice site mutations in genes that cause hundreds of diseases.
Split gene theory requires a separate origin of all eukaryotic species. It also requires that the simpler prokaryotes evolved from eukaryotes. This completely contradicts the scientific consensus about the formation of eukaryotic cells by endosymbiosis of bacteria. In 1994, Senapathy wrote a book about this aspect of his theory - The Independent Birth of Organisms. It proposed that all eukaryotic genomes were formed separately in a primordial pool. Dutch biologist Gert Korthoff criticized the theory by posing various problems that cannot be explained by a theory of independent origins. He pointed out that various eukaryotes need nurturing and called this the 'boot problem', in that even the initial eukaryote needed parental care. Korthoff notes that a large fraction of eukaryotes are parasites. Senapathy's theory would require a coincidence to explain their existence.[4][5] Senapathy's theory cannot explain the strong evidence for common descent (homology, universal genetic code, embryology, fossil record.)[6]
Background
[edit]
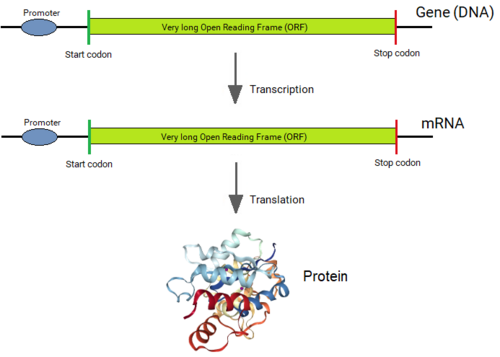
Genes of all organisms, except bacteria, consist of short protein-coding regions (exons) interrupted by long sequences (introns).[1][2] When a gene is expressed, its DNA sequence is copied into a “primary RNA” sequence by the enzyme RNA polymerase. Then the “spliceosome” machinery physically removes the introns from the RNA copy of the gene by the process of splicing, leaving only a contiguously connected series of exons, which becomes messenger RNA (mRNA). This mRNA is now read by the ribosome, which produces the encoded protein. Thus, although introns are not physically removed from a gene, a gene's sequence is read as if introns were not present.
Exons are usually short, with an average length of about 120 bases (e.g. in human genes). Intron lengths vary widely from 10 to 500,000, but exon lengths have an upper bound of about 600 bases in most eukaryotes. Because exons code for protein sequences, they are important for the cell, yet constitute only ~2% of the sequences. Introns, in contrast, constitute 98% of the sequences but seem to have few crucial functions, except for enhancer sequences and developmental regulators in rare instances.[7][8]
Until Philip Sharp[9][10] and Richard Roberts[11] discovered introns[12] within eukaryotic genes in 1977, it was believed that the coding sequence of all genes was always in one single stretch, bounded by a single long ORF. The discovery of introns was a profound surprise, which instantly brought up the questions of how, why and when the introns came into the eukaryotic genes.
It soon became apparent that a typical eukaryotic gene was interrupted at many locations by introns, dividing the coding sequence into many short exons. Also surprising was that the introns were long, as long as hundreds of thousands of bases. These findings prompted the questions of why many introns occur within a gene (for example, ~312 introns occur in the human gene TTN), why they are long, and why exons are short.
| Gene symbol | Gene length (bases) |
Longest Intron length (bases) |
Number of
introns in the gene |
|---|---|---|---|
| ROBO2 | 1,743,269 | 1,160,411 | 104 |
| KCNIP4 | 1,220,183 | 1,097,903 | 76 |
| ASIC2 | 1,161,877 | 1,043,911 | 18 |
| NRG1 | 1,128,573 | 956,398 | 177 |
| DPP10 | 1,403,453 | 866,399 | 142 |
It was also discovered that the spliceosome machinery was large and complex with ~300 proteins and several SnRNA molecules. The questions extended to the origin of the spliceosome. Soon after the discovery of introns, it became apparent that the junctions between exons and introns on either side exhibited specific sequences that directed the spliceosome machinery to the exact base position for splicing. How and why these splice junction signals came into being was another important question.
History
[edit]The discovery of introns and the split gene architecture of the eukaryotic genes started a new era of eukaryotic biology. The question of why eukaryotic genes had fragmented genes prompted speculation and discussion almost immediately.
Ford Doolittle published a paper in 1978 in which he stated that most molecular biologists assumed that the eukaryotic genome arose from a ‘simpler’ and more ‘primitive’ prokaryotic genome rather like that of Escherichia coli.[13] However, this type of evolution would require that introns be introduced into the coding sequences of bacterial genes. Regarding this requirement, Doolittle said, “It is extraordinarily difficult to imagine how informationally irrelevant sequences could be introduced into pre-existing structural genes without deleterious effects.” He stated “I would like to argue that the eukaryotic genome, at least in that aspect of its structure manifested as ‘genes in pieces’ is in fact the primitive original form.”
James Darnell expressed similar views in 1978. He stated, “The differences in the biochemistry of messenger RNA formation in eukaryotes compared to prokaryotes are so profound as to suggest that sequential prokaryotic to eukaryotic cell evolution seems unlikely. The recently discovered non-contiguous sequences in eukaryotic DNA that encode messenger RNA may reflect an ancient, rather than a new, distribution of information in DNA and that eukaryotes evolved independently of prokaryotes.”[14]
However, in an apparent attempt to reconcile with the idea that RNA preceded DNA in evolution, and with the concept of the three evolutionary lineages of archea, bacteria and eukarya, both Doolittle and Darnell deviated from their original speculation in a joint paper in 1985.[15] They suggested that the ancestor of all three groups of organisms, the ‘progenote,’ had a genes-in-pieces structure, from which all three lineages evolved. They speculated that the precellular stage had primitive RNA genes which had introns, which were reverse transcribed into DNA and formed the progenote. Bacteria and archea evolved from the progenote by losing introns, and ‘urkaryote’ evolved from it by retaining introns. Later, the eukaryote evolved from the urkaryote by evolving a nucleus and absorbing mitochondria from bacteria. Multicellular organisms then evolved from the eukaryote.
These authors predicted that the distinctions between the prokaryote and the eukaryote were so profound that the prokaryote to eukaryote evolution was not tenable, and had different origins. However, other than the speculations that the precellular RNA genes must have had introns, they did not address the key questions of intron origin. No explanations described why exons were short and introns were long, how the splice junctions originated, what the structure and sequence of the splice junctions meant, and why eukaryote genomes were large.
Around the same time that Doolittle and Darnell suggested that introns in eukaryotic genes could be ancient, Colin Blake[16] and Walter Gilbert[17][18] published their views on intron origins independently. In their view, introns originated as spacer sequences that enabled convenient recombination and shuffling of exons that encoded distinct functional domains in order to evolve new genes. Thus, new genes were assembled from exon modules that coded for functional domains, folding regions, or structural elements from preexisting genes in the genome of an ancestral organism, thereby evolving genes with new functions. They did not specify how exons or introns originated. In addition, even after many years, extensive analysis of thousands of proteins and genes showed that only extremely rarely do genes exhibit the supposed exon shuffling phenomenon.[19][20] Furthermore, molecular biologists questioned the exon shuffling proposal, from a purely evolutionary view for both methodological and conceptual reasons, and, in the long run, this theory did not survive.
Hypothesis
[edit]Around the time introns were discovered, Senapathy was asking how genes themselves could have originated. He surmised that for any gene to come into being, genetic sequences (RNA or DNA) must have been present in the prebiotic environment. A basic question he asked was how protein-coding sequences could have originated from primordial DNA sequences at the origin of the first cells.
To answer this, he made two basic assumptions:
- before a self-replicating cell could come into existence, DNA molecules were synthesized in the primordial soup by random addition of the 4 nucleotides without the help of templates and
- the nucleotide sequences that code for proteins were selected from these preexisting random DNA sequences in the primordial soup, and not by construction from shorter coding sequences.
He also surmised that codons must have been established prior to the origin of the first genes. If primordial DNA did contain random nucleotide sequences, he asked: Was there an upper limit in coding-sequence lengths, and, if so, did this limit play a crucial role in the formation of the structural features of genes at the origin of genes?
His logic was the following. The average length of proteins in living organisms, including the eukaryotic and bacterial organisms, was ~400 amino acids. However, much longer proteins existed, even longer than 10,000-30,000 amino acids in both eukaryotes and bacteria.[21] Thus, the coding sequence of thousands of bases existed in a single stretch in bacterial genes. In contrast, the coding sequence of eukaryotes existed only in short segments of exons of ~120 bases regardless of the length of the protein. If the coding sequence ORF lengths in random DNA sequences were as long as those in bacterial organisms, then long, contiguous coding genes were possible in random DNA. This was not known, as the distribution of ORF lengths in a random DNA sequence had never been studied.
As random DNA sequences could be generated in the computer, Senapathy thought that he could ask these questions and conduct his experiments in silico. Furthermore, when he began studying this question, sufficient DNA and protein sequence information existed in the National Biomedical Research Foundation (NBRF) database in the early 1980s.
Testing the hypothesis
[edit]Origin of introns/split genes
[edit]
Senapathy analyzed the distribution of the ORF lengths in computer-generated random DNA sequences first. Surprisingly, this study revealed that about 200 codons (600 bases) was the upper limit in ORF lengths. The shortest ORF (zero base in length) was the most frequent. At increasing lengths of ORFs, their frequency decreased logarithmically, approaching zero at about 600 bases. When the probability of ORF lengths in a random sequence was plotted, it revealed that the probability of increasing lengths of ORFs decreased exponentially and tailed off at a maximum of about 600 bases. From this “negative exponential” distribution of ORF lengths, it was found that most of ORFs were far shorter than the maximum.

This finding was surprising because the coding sequence for the average protein length of 400 AAs (with ~1,200 bases of coding sequence) and longer proteins of thousands of AAs (requiring >10,000 bases of coding sequence) would not occur at a stretch in a random sequence. If this was true, a typical gene with a contiguous coding sequence could not originate in a random sequence. Thus, the only possible way that any gene could originate from a random sequence was to split the coding sequence into shorter segments and select these segments from short ORFs available in the random sequence, rather than to increase the ORF length by eliminating consecutive stop codons. This process of choosing short segments of coding sequences from the available ORFs to make a long ORF would lead to a split structure.
If this hypothesis was true, eukaryotic DNA sequences should reflect it. When Senapathy plotted the distribution of ORF lengths in eukaryotic DNA sequences, the plot was remarkably similar to that from random DNA sequences. This plot was also a negative exponential distribution that tailed off at a maximum of about 600 bases, as with eukaryotic genes,[1][22][3] which coincided exactly with the maximum length of ORFs observed in both random DNA and eukaryotic DNA sequences.
The split genes thus originated from random DNA sequences by choosing the best of the short coding segments (exons) and splicing them. The intervening intron sequences were left-over vestiges of the random sequences, and thus were earmarked to be removed by the spliceosome. These findings indicated that split genes could have originated from random DNA sequences with exons and introns as they appear in today's eukaryotic organisms. Nobel Laureate Marshall Nirenberg, who deciphered the codons, stated that these findings strongly showed that the split gene theory for the origin of introns and the split structure of genes must be valid.[1][23]
Blake proposed the Gilbert-Blake hypothesis in 1979 for the origin of introns and stated that Senapathy's split gene theory comprehensively explained the origin of the split gene structure. In addition, he stated that it explained several key questions including the origin of the splicing mechanism:[16]
Recent work by Senapathy, when applied to RNA, comprehensively explains the origin of the segregated form of RNA into coding and non-coding regions. It also suggests why a splicing mechanism was developed at the start of primordial evolution. He found that the distribution of reading frame lengths in a random nucleotide sequence corresponded exactly to that for the observed distribution of eukaryotic exon sizes. These were delimited by regions containing stop signals, the messages to terminate construction of the polypeptide chain, and were thus non-coding regions or introns. The presence of a random sequence was therefore sufficient to create in the primordial ancestor the segregated form of RNA observed in the eukaryotic gene structure. Moreover, the random distribution also displays a cutoff at 600 nucleotides, which suggests that the maximum size for an early polypeptide was 200 residues, again as observed in the maximum size of the eukaryotic exon. Thus, in response to evolutionary pressures to create larger and more complex genes, the RNA fragments were joined together by a splicing mechanism that removed the introns. Hence, the early existence of both introns and RNA splicing in eukaryotes appears to be very likely from a simple statistical basis. These results also agree with the linear relationship found between the number of exons in the gene for a particular protein and the length of the polypeptide chain.”

Origin of splice junctions
[edit]
Under the split gene theory, an exon is defined by an ORF. It requires a mechanism to recognize an ORF to have originated. As an ORF is defined by a contiguous coding sequence bounded by stop codons, these stop codon ends had to be recognized by the exon-intron gene recognition system. This system could have defined the exons by the presence of a stop codon at the ends of ORFs, which should be included within the ends of the introns and eliminated by the splicing process. Thus, the introns should contain a stop codon at their ends, which would be part of the splice junction sequences.
If this hypothesis was true, the split genes of today's living organisms should contain stop codons exactly at the ends of introns. When Senapathy tested this hypothesis in the splice junctions of eukaryotic genes, he found that the vast majority of splice junctions did contain a stop codon at the end of each intron, outside of the exons. In fact, these stop codons were found to form the “canonical” GT:AG splicing sequence, with the three stop codons occurring as part of the strong consensus signals. Thus, the basic split gene theory for the origin of introns and the split gene structure led to the understanding that the splice junctions originated from the stop codons.[2]
| Codon | Number of occurrences in donor signal |
Number of occurrences in acceptor signal |
|---|---|---|
| TAA | 370 | 0 |
| TGA | 293 | 0 |
| TAG | 64 | 234 |
| CAG | 7 | 746 |
| Other | 297* | 50 |
| Total | 1030 | 1030 |
| *More than 70% are TAX [TAT = 75; TAC = 59; TGT = 70]. | ||
Sequence data for only about 1,000 exon-intron junctions were available when Senapathy thought about this question. He took the data for 1,030 splice junction sequences (donors and acceptors) and counted the codons occurring at each of the 7- base positions in the donor signal sequence [CAG:GTGAGT] and each of the possible 2-base positions in the acceptor signal [CAG:G] from the GenBank database. He found that the stop codons occurred at high frequency only at the 5th base position in the donor signal and the first base position in the acceptor signal. These positions are the* start of the intron (in fact, one base after the start) and at the end of the intron, as Senapathy had predicted. The codon counts at only these positions are shown. Even when the codons at these positions were not stop codons, 70% of them began with the first two bases of the stop codons TA and TG [TAT = 75; TAC = 59; TGT = 70].
All three stop codons (TGA, TAA and TAG) were found after one base (G) at the start of introns. These stop codons are shown in the consensus canonical donor splice junction as AG:GT(A/G)GGT, wherein the TAA and TGA are the stop codons, and the additional TAG is also present at this position. Besides the codon CAG, only TAG, which is a stop codon, was found at the ends of introns. The canonical acceptor splice junction is shown as (C/T)AG:GT, in which TAG is the stop codon. These consensus sequences clearly show the presence of the stop codons at the ends of introns bordering the exons in all eukaryotic genes, thus providing a strong corroboration for the split gene theory. Nirenberg again stated that these observations fully supported the split gene theory for the origin of splice junction sequences from stop codons.[2][24]
Soon after the discovery of introns by Philip Sharp and Richard Roberts, it became known that mutations within splice junctions could lead to diseases. Senapathy showed that mutations in the stop codon bases (canonical bases) caused more diseases than the mutations in non-canonical bases.[1]
Branch point (lariat) sequence
[edit]An intermediate stage in the process of eukaryotic RNA splicing is the formation of a lariat structure. It is anchored at an adenosine residue in intron between 10 and 50 nucleotides upstream of the 3' splice site. A short conserved sequence (the branch point sequence) functions as the recognition signal for the site of lariat formation. During the splicing process, this conserved sequence towards the end of the intron forms a lariat structure with the beginning of the intron.[25] The final step of the splicing process occurs when the two exons are joined and the intron is released as a lariat RNA.[26]
Several investigators found the branch point sequences in different organisms[25] including yeast, human, fruit fly, rat, and plants. Senapathy found that, in all of these sequences, the codon ending at the branch point adenosine is consistently a stop codon. What is interesting is that two of the three stop codons (TAA and TGA) occur almost all of the time at this position.
| Organism | Lariat Consensus sequence |
|---|---|
| Yeast | TACTAAC |
| Human Beta globin genes | CTGAC
CTAAT CTGAT CTAAC CTCAC |
| Drosophila | CTAAT |
| Rats | CTGAC |
| Plants | (C/T)T(A/G)A(T/C) |
| Consistent presence of stop codons in branch point signal sequences.
Lariat (branch point) sequences have been identified from many different organisms.These sequences consistently show that the codon ending in the branching adenosine is a stop codon, either TAA or TGA, which are shown in red. | |
These findings led Senapathy to propose that the branch point signal originated from stop codons. The finding that two different stop codons (TAA and TGA) occur within the lariat signal with the branching point as the third base of the stop codons corroborates this proposal. As the branching point of the lariat occurs at the last adenine of the stop codon, it is possible that the spliceosome machinery that originated for the elimination of the stop codons from the primary RNA sequence created an auxiliary stop-codon sequence signal as the lariat sequence to aid its splicing function.[2]
The small nuclear U2 RNA found in splicing complexes is thought to aid splicing by interacting with the lariat sequence.[27] Complementary sequences for both the lariat sequence and the acceptor signal are present in a segment of only 15 nucleotides in U2 RNA. Further, the U1 RNA has been proposed to function as a guide in splicing to identify the precise donor splice junction by complementary base-pairing. The conserved regions of the U1 RNA thus include sequences complementary to the stop codons. These observations enabled Senapathy to predict that stop codons had operated in the origin of not only the splice-junction signals and the lariat signal, but also some small nuclear RNAs.
Gene regulatory sequences
[edit]Senapathy proposed that the gene-expression regulatory sequences (promoter and poly-A addition site sequences) also could have originated from stop codons. A conserved sequence, AATAAA, exists in almost every gene a short distance downstream from the end of the protein-coding message and serves as a signal for the addition of poly(A) in the mRNA copy of the gene.[28] This poly(A) sequence signal contains a stop codon, TAA. A sequence shortly downstream from this signal, thought to be part of the complete poly(A) signal, also contains the TAG and TGA stop codons.
Eukaryotic RNA-polymerase-II-dependent promoters can contain a TATA box (consensus sequence TATAAA), which contains the stop codon TAA. Bacterial promoter elements at ~10 bases exhibits a TATA box with a consensus of TATAAT (which contains the stop codon TAA), and at -35 bases exhibits a consensus of TTGACA (containing the stop codon TGA). Thus, the evolution of the whole RNA processing mechanism seems to have been influenced by the too-frequent occurrence of stop codons, thus making the stop codons the focal points for RNA processing.
Stop codons are key parts of every genetic element in the eukaryotic gene
[edit]
| Genetic Element | Consensus sequence |
|---|---|
| Promoter | TATAAT |
| Donor Splice Sequence | CAG:GTAAGT
CAG:GTGAGT |
| Acceptor Splice Sequence | (C/T)9...TAG:GT |
| Lariat Sequence | CTGAC
CTAAC |
| Poly-A addition site | TATAAA |
| The consistent occurrence of stop codons in genetic elements in eukaryotic genes.The consensus sequences of the different genetic elements in eukaryotic genes are shown. The stop codon(s) in each of these sequences are colored in red. | |
Senapathy discovered that stop codons occur as key parts in every genetic element in eukaryotic genes. The table and figure show that the key parts of the core promoter elements, the lariat signal, the donor and acceptor splice signals, and the poly-A addition signal consist of one or more stop codons. This finding corroborates the split gene theory's claim that the underlying reason for the complete split gene paradigm is the origin of split genes from random DNA sequences, wherein random distribution of an extremely high frequency of stop codons were used by nature to define these genetic elements.
Short exons/long introns
[edit]Research based on the split gene theory sheds light on other basic questions of exons and introns. The exons of eukaryotes are generally short (human exons average ~120 bases, and can be as short as 10 bases) and introns are usually long (average of ~3,000 bases, and can be several hundred thousands bases long), for example genes RBFOX1, CNTNAP2, PTPRD and DLG2. Senapathy provided a plausible answer to these questions, the only explanation to date. If eukaryotic genes originated from random DNA sequences, they have to match the lengths of ORFs from random sequences, and possibly should be around 100 bases (close to the median length of ORFs in random sequence). The genome sequences of living organisms exhibit exactly the same average lengths of 120 bases for exons, and the longest exons of 600 bases (with few exceptions), which is the same length as that of the longest random ORFs.[1][2][3][22]
If split genes originated in random DNA sequences, then introns would be long for several reasons. The stop codons occur in clusters leading to numerous consecutive short ORFs: longer ORFs that could be defined as exons would be rarer. Furthermore, the best of the coding sequence parameters for functional proteins would be chosen from the long ORFs in random sequence, which may occur rarely. In addition, the combination of donor and acceptor splice junction sequences within short lengths of coding sequence segments that would define exon boundaries would occur rarely in a random sequence. These combined reasons would make introns long compared to exons.
Eukaryotic genomes
[edit]This work also explains why genomes such as the human genome have billions of bases, and why only a small fraction (~2%) codes for proteins and other regulatory elements.[29][30] If split genes originated from random primordial DNA sequences, they would contain a significant amount of DNA that represented by introns. Furthermore, a genome assembled from random DNA containing split genes would also include intergenic random DNA. Thus, genomes that originated from random DNA sequences had to be large, regardless of the complexity of the organism.
The observation that several organisms such as the onion (~16 billion bases[31]) and salamander (~32 billion bases[32]) have much larger genomes than humans (~3 billion bases[33][34]) while the organisms are no more complex than humans comports with the theory. Furthermore, the fact that several organisms with smaller genomes have a similar number of genes as human, such as C. elegans (genome size ~100 million bases, ~19,000 genes)[35] and Arabidopsis thaliana (genome size ~125 million bases, ~25,000 genes),[36] supports the theory. The theory predicts that the introns in the split genes in these genomes could be the “reduced” (or deleted) form compared to larger genes with long introns, thus leading to reduced genomes.[1][22] In fact, researchers have recently proposed that these smaller genomes are actually reduced genomes.[37]
Spliceosomal machinery and eukaryotic nucleus
[edit]Senapathy addressed the origin of the spliceosomal machinery that edits out the introns from RNA transcripts. If the split genes had originated from random DNA, then the introns would have become an unnecessary but integral part of eukaryotic genes along with the splice junctions. The spliceosomal machinery would be required to remove them and to enable the short exons to be linearly spliced together as a contiguously coding mRNA that can be translated into a complete protein. Thus, the split gene theory argues that spliceosomal machinery exists to remove the unnecessary introns.[1][2]
Blake states, “Work by Senapathy, when applied to RNA, comprehensively explains the origin of the segregated form of RNA into coding and noncoding regions. It also suggests why a splicing mechanism was developed at the start of primordial evolution.”[16]
Eukaryotes
[edit]Senapathy proposed a plausible mechanistic and functional rationale why the eukaryotic nucleus originated, a major question in biology.[1][2] If the transcripts of the split genes and the spliced mRNAs were present in a cell without a nucleus, the ribosomes would try to bind to both the un-spliced primary RNA transcript and the spliced mRNA, which would result in chaos. A boundary that separates the RNA splicing process from the mRNA translation avoids this problem. The nuclear boundary provides a clear separation of the primary RNA splicing and the mRNA translation.
These investigations thus led to the possibility that primordial DNA with essentially random sequence gave rise to the complex structure of the split genes with exons, introns and splice junctions. Cells that harbored split genes had to be complex with a nuclear cytoplasmic boundary, and must have a spliceosomal machinery. Thus, it was possible that the earliest cell was complex and eukaryotic.[1][2][3][22] Surprisingly, findings from extensive comparative genomics research from several organisms since 2007 overwhelmingly show that the earliest organisms could have been highly complex and eukaryotic, and could have contained complex proteins,[38][39][40][41][42][43][44] as predicted by Senapathy's theory.
The spliceosome is a highly complex mechanism, containing ~200 proteins and several SnRNPs. Collins and Penny stated, “We begin with the hypothesis that ... the spliceosome has increased in complexity throughout eukaryotic evolution. However, examination of the distribution of spliceosomal components indicates that not only was a spliceosome present in the eukaryotic ancestor but it also contained most of the key components found in today's eukaryotes. ... the last common ancestor of extant eukaryotes appears to show much of the molecular complexity seen today.” This suggests that the earliest eukaryotic organisms were complex and contained sophisticated genes and proteins.[45]
Bacterial genes
[edit]Genes with uninterrupted coding sequences that are thousands of bases long - up to 90,000 bases - that occur in many bacterial organisms[21] were practically impossible to have occurred. However, the bacterial genes could have originated from split genes by losing introns, the only proposed way to arrive at long coding sequences. It is also a better[clarification needed] way than by increasing the lengths of ORFs from short random ORFs to long ORFs by specifically removing the stop codons by mutation.[1][2][3]
| Gene size (bases) | Number of genes |
|---|---|
| 5,000 - 10,000 | 3,029 |
| 10,000 - 15,000 | 492 |
| 15,000 - 20,000 | 131 |
| 20,000 - 25,000 | 39 |
| >25,000 | 41 |
| Long coding sequences occur as long ORFs in bacterial genes. Thousands of genes that are longer than 5,000 bases, coding for proteins that are longer than 2,000 amino acids, exist in many bacterial genomes. The longest protein coding genes are ~90,000 bases long. Each occurs in a single stretch of coding sequence (ORF) without stop codons or introns.[21] | |
According to the split gene theory, this process of intron loss could have happened from prebiotic random DNA. These contiguously coding genes could be tightly organized in the bacterial genomes without any introns and be more streamlined. According to Senapathy, the nuclear boundary that was required for a cell containing split genes would not be required for a cell containing only uninterrupted genes. Thus, the bacterial cells did not develop a nucleus. Based on split gene theory, the eukaryotic genomes and bacterial genomes could have independently originated from the split genes in primordial random DNA sequences.
Shapiro-Senapathy algorithm
[edit]Senapathy developed algorithms to detect donor and acceptor splice sites, exons and a complete split gene in a genomic sequence. He developed the position weight matrix (PWM) method based on the frequency of the four bases at the consensus sequences of the donor and acceptor in different organisms to identify the splice sites in a given sequence. Furthermore, he formulated the first algorithm to find the exons based on the requirement of exons to contain a donor sequence (at the 5’ end) and an acceptor sequence (at the 3’ end), and an ORF in which the exon should occur, and another algorithm to find a complete split gene. These algorithms are collectively known as the Shapiro-Senapathy algorithm (S&S).[46][47]
This algorithm aids in the identification of splicing mutations that cause disease and adverse drug reactions.[46][47] Scientists used the algorithm to identify mutations and genes that cause cancers, inherited disorders, immune deficiency diseases and neurological disorders. It is increasingly used in clinical practice and research to find mutations in known disease-causing genes in patients and to discover novel genes that are causal of different diseases. Furthermore, it is used in defining the cryptic splice sites and deducing the mechanisms by which mutations can affect normal splicing and lead to different diseases. It is also employed in basic research.
Findings based on S&S have impacted major questions in eukaryotic biology and in human medicine.[48]
Corroborating evidence
[edit]The split gene theory implies that structural features of split genes predicted from computer-simulated random sequences occur in eukaryotic split genes. This is borne out in most known split genes. The sequences exhibit a nearly perfect negative exponential distribution of ORF lengths.[1][2][22][3] With rare exceptions, eukaryotic gene exons fall within the predicted 600 base maximum.
The theory correctly predicts that exons are delimited by stop codons, especially at the 3’ ends of exons. Actually they are precisely delimited more strongly at the 3’ ends of exons and less strongly at the 5’ ends in most known genes, as predicted.[1][2][22][3] These stop codons are the most important functional parts of both splice junctions. The theory thus provides an explanation for the “conserved” splice junctions at the ends of exons and for the loss of these stop codons along with introns when they are spliced out. The theory correctly predicts that splice junctions are randomly distributed in eukaryotic DNA sequences.[3][25][46][47] The theory correctly predicts that splice junctions present in transfer RNA genes and ribosomal RNA genes, do not contain stop codons. The lariat signal, another sequence involved in the splicing process, also contains stop codons.[1][2][3][22][25][46][47]
The theory correctly predicts that introns are non-coding and that they are mostly non-functional. Except for some intron sequences including the donor and acceptor splice signal sequences and branch point sequences, and possibly the intron splice enhancers that occur at the ends of introns, which aid in the removal of introns, the vast majority of introns are devoid of any functions. The theory does not exclude rare sequences within introns that could be used by the genome and the cell, especially because introns are so long.
Thus, the theory's predictions are precisely corroborated by the major elements in modern eukaryotic genomes.
Comparative analysis of the modern genome data from several living organisms found that the characteristics of split genes trace back to the earliest organisms. These organisms could have contained the split genes and complex proteins that occur in today's living organisms.[49][50][51][52][53][54][55][56][57]
Studies employing maximum likelihood analysis found that the earliest eukaryotic organisms contained the same genes as modern organisms with yet a higher intron density.[58] Comparative genomics of many organisms including basal eukaryotes[59] (considered to be primitive eukaryotic organisms such as Amoeboflagellata, Diplomonadida, and Parabasalia) showed that intron-rich split genes accompanied and spliceosome from modern organisms were present in their earliest forebears, and that the earliest organisms came with all the eukaryotic cellular components.[60][49][61][62][63][58]
Selected publications
[edit]- Shapiro, Marvin B.; Senapathy, Periannan (1987). "RNA splice junctions of different classes of eukaryotes: sequence statistics and functional implications in gene expression". Nucleic Acids Research. 15 (17): 7155–7174. doi:10.1093/nar/15.17.7155. PMC 306199. PMID 3658675.
- Senapathy, P. (1988). "Possible evolution of splice-junction signals in eukaryotic genes from stop codons". Proc Natl Acad Sci U S A. 85 (4): 1129–33. Bibcode:1988PNAS...85.1129S. doi:10.1073/pnas.85.4.1129. PMC 279719. PMID 3422483.
- Senapathy, P; Shapiro, MB; Harris, NL (1990). [16] Splice junctions, branch point sites, and exons: Sequence statistics, identification, and applications to genome project. Methods in Enzymology. Vol. 183. pp. 252–78. doi:10.1016/0076-6879(90)83018-5. ISBN 9780121820848. PMID 2314278.
- Harris, N.L.; Senapathy, P. (1990). "Distribution and consensus of branch point signals in eukaryotic genes: a computerized statistical analysis". Nucleic Acids Res. 18 (10): 3015–9. doi:10.1093/nar/18.10.3015. PMC 330832. PMID 2349097.
- Senapathy, P. (1986). "Origin of eukaryotic introns: a hypothesis, based on codon distribution statistics in genes, and its implications". Proc Natl Acad Sci U S A. 83 (7): 2133–7. Bibcode:1986PNAS...83.2133S. doi:10.1073/pnas.83.7.2133. PMC 323245. PMID 3457379.
- Regulapati, R.; Bhasi, A.; Singh, C.K.; Senapathy, P. (2008). "Origination of the Split Structure of Spliceosomal Genes from Random Genetic Sequences". PLOS ONE. 3 (10): 10. Bibcode:2008PLoSO...3.3456R. doi:10.1371/journal.pone.0003456. PMC 2565106. PMID 18941625.
- Senapathy, P. (1995). "Introns and the origin of protein-coding genes". Science. 268 (5215): 1366–7. Bibcode:1995Sci...268.1366S. doi:10.1126/science.7761858. PMID 7761858.
References
[edit]- ^ a b c d e f g h i j k l m n o p q Senapathy, P. (April 1986). "Origin of eukaryotic introns: a hypothesis, based on codon distribution statistics in genes, and its implications". Proceedings of the National Academy of Sciences of the United States of America. 83 (7): 2133–2137. Bibcode:1986PNAS...83.2133S. doi:10.1073/pnas.83.7.2133. ISSN 0027-8424. PMC 323245. PMID 3457379.
- ^ a b c d e f g h i j k l m n o Senapathy, P. (February 1982). "Possible evolution of splice-junction signals in eukaryotic genes from stop codons". Proceedings of the National Academy of Sciences of the United States of America. 85 (4): 1129–1133. Bibcode:1988PNAS...85.1129S. doi:10.1073/pnas.85.4.1129. ISSN 0027-8424. PMC 279719. PMID 3422483.
- ^ a b c d e f g h i j Senapathy, P. (1995-06-02). "Introns and the origin of protein-coding genes". Science. 268 (5215): 1366–1367, author reply 1367–1369. Bibcode:1995Sci...268.1366S. doi:10.1126/science.7761858. ISSN 0036-8075. PMID 7761858.
- ^ "Independent origin and the facts of life". wasdarwinwrong.com. Retrieved 2021-07-31.
- ^ "Independent Birth of Organisms. Periannan Senapathy. Book review". wasdarwinwrong.com. Retrieved 2021-07-31.
- ^ Theobald, Douglas L. (2012). "29+ Evidences for Macroevolution: The Scientific Case for Common Descent".
{{cite journal}}: Cite journal requires|journal=(help) - ^ Gillies, S. D.; Morrison, S. L.; Oi, V. T.; Tonegawa, S. (June 1983). "A tissue-specific transcription enhancer element is located in the major intron of a rearranged immunoglobulin heavy chain gene". Cell. 33 (3): 717–728. doi:10.1016/0092-8674(83)90014-4. ISSN 0092-8674. PMID 6409417. S2CID 40313833.
- ^ Mercola, M.; Wang, X. F.; Olsen, J.; Calame, K. (1983-08-12). "Transcriptional enhancer elements in the mouse immunoglobulin heavy chain locus". Science. 221 (4611): 663–665. Bibcode:1983Sci...221..663M. doi:10.1126/science.6306772. ISSN 0036-8075. PMID 6306772.
- ^ Berk, A. J.; Sharp, P. A. (November 1977). "Sizing and mapping of early adenovirus mRNAs by gel electrophoresis of S1 endonuclease-digested hybrids". Cell. 12 (3): 721–732. doi:10.1016/0092-8674(77)90272-0. ISSN 0092-8674. PMID 922889.
- ^ Berget, S M; Moore, C; Sharp, P A (August 1977). "Spliced segments at the 5' terminus of adenovirus 2 late mRNA". Proceedings of the National Academy of Sciences of the United States of America. 74 (8): 3171–3175. Bibcode:1977PNAS...74.3171B. doi:10.1073/pnas.74.8.3171. ISSN 0027-8424. PMC 431482. PMID 269380.
- ^ Chow, L. T.; Roberts, J. M.; Lewis, J. B.; Broker, T. R. (August 1977). "A map of cytoplasmic RNA transcripts from lytic adenovirus type 2, determined by electron microscopy of RNA:DNA hybrids". Cell. 11 (4): 819–836. doi:10.1016/0092-8674(77)90294-X. ISSN 0092-8674. PMID 890740. S2CID 37967144.
- ^ "Online Education Kit: 1977: Introns Discovered". National Human Genome Research Institute (NHGRI). Retrieved 2019-01-01.
- ^ Doolittle, W. Ford (13 April 1978). "Genes in pieces: were they ever together?". Nature. 272 (5654): 581–582. Bibcode:1978Natur.272..581D. doi:10.1038/272581a0. ISSN 1476-4687. S2CID 4162765.
- ^ Darnell, J. E. (1978-12-22). "Implications of RNA-RNA splicing in evolution of eukaryotic cells". Science. 202 (4374): 1257–1260. doi:10.1126/science.364651. ISSN 0036-8075. PMID 364651.
- ^ Doolittle, W. F.; Darnell, J. E. (1986-03-01). "Speculations on the early course of evolution". Proceedings of the National Academy of Sciences. 83 (5): 1271–1275. Bibcode:1986PNAS...83.1271D. doi:10.1073/pnas.83.5.1271. ISSN 1091-6490. PMC 323057. PMID 2419905.
- ^ a b c Blake, C.C.F. (1985-01-01). Exons and the Evolution of Proteins. International Review of Cytology. Vol. 93. pp. 149–185. doi:10.1016/S0074-7696(08)61374-1. ISBN 9780123644930. ISSN 0074-7696. PMID 2409042.
- ^ Gilbert, Walter (February 1978). "Why genes in pieces?". Nature. 271 (5645): 501. Bibcode:1978Natur.271..501G. doi:10.1038/271501a0. ISSN 1476-4687. PMID 622185. S2CID 4216649.
- ^ Tonegawa, S; Maxam, A M; Tizard, R; Bernard, O; Gilbert, W (March 1978). "Sequence of a mouse germ-line gene for a variable region of an immunoglobulin light chain". Proceedings of the National Academy of Sciences of the United States of America. 75 (3): 1485–1489. Bibcode:1978PNAS...75.1485T. doi:10.1073/pnas.75.3.1485. ISSN 0027-8424. PMC 411497. PMID 418414.
- ^ Feng, D. F.; Doolittle, R. F. (1987-01-01). "Reconstructing the Evolution of Vertebrate Blood Coagulation from a Consideration of the Amino Acid Sequences of Clotting Proteins". Cold Spring Harbor Symposia on Quantitative Biology. 52: 869–874. doi:10.1101/SQB.1987.052.01.095. ISSN 1943-4456. PMID 3483343.
- ^ Gibbons, A. (1990-12-07). "Calculating the original family--of exons". Science. 250 (4986): 1342. Bibcode:1990Sci...250.1342G. doi:10.1126/science.1701567. ISSN 1095-9203. PMID 1701567.
- ^ a b c Reva, Oleg; Tümmler, Burkhard (2008). "Think big – giant genes in bacteria" (PDF). Environmental Microbiology. 10 (3): 768–777. Bibcode:2008EnvMi..10..768R. doi:10.1111/j.1462-2920.2007.01500.x. hdl:2263/9009. ISSN 1462-2920. PMID 18237309.
- ^ a b c d e f g Regulapati, Rahul; Singh, Chandan Kumar; Bhasi, Ashwini; Senapathy, Periannan (2008-10-20). "Origination of the Split Structure of Spliceosomal Genes from Random Genetic Sequences". PLOS ONE. 3 (10): e3456. Bibcode:2008PLoSO...3.3456R. doi:10.1371/journal.pone.0003456. ISSN 1932-6203. PMC 2565106. PMID 18941625.
- ^ New Scientist. Reed Business Information. 1986-06-26.
- ^ New Scientist. Reed Business Information. 1988-03-31.
- ^ a b c d Senapathy, Periannan; Harris, Nomi L. (1990-05-25). "Distribution and consenus of branch point signals in eukaryotic genes: a computerized statistical analysis". Nucleic Acids Research. 18 (10): 3015–9. doi:10.1093/nar/18.10.3015. ISSN 0305-1048. PMC 330832. PMID 2349097.
- ^ Maier, U.-G.; Brown, J.W.S.; Toloczyki, C.; Feix, G. (January 1987). "Binding of a nuclear factor to a consensus sequence in the 5' flanking region of zein genes from maize". The EMBO Journal. 6 (1): 17–22. doi:10.1002/j.1460-2075.1987.tb04712.x. ISSN 0261-4189. PMC 553350. PMID 15981330.
- ^ Keller, E B; Noon, W A (1985-07-11). "Intron splicing: a conserved internal signal in introns of Drosophila pre-mRNAs". Nucleic Acids Research. 13 (13): 4971–4981. doi:10.1093/nar/13.13.4971. ISSN 0305-1048. PMC 321838. PMID 2410858.
- ^ BIRNSTIEL, M; BUSSLINGER, M; STRUB, K (June 1985). "Transcription termination and 3′ processing: the end is in site!". Cell. 41 (2): 349–359. doi:10.1016/s0092-8674(85)80007-6. ISSN 0092-8674. PMID 2580642. S2CID 11999043.
- ^ Consortium, International Human Genome Sequencing (February 2001). "Initial sequencing and analysis of the human genome". Nature. 409 (6822): 860–921. Bibcode:2001Natur.409..860L. doi:10.1038/35057062. hdl:2027.42/62798. ISSN 1476-4687. PMID 11237011.
- ^ Zhu, Xiaohong; Zandieh, Ali; Xia, Ashley; Wu, Mitchell; Wu, David; Wen, Meiyuan; Wang, Mei; Venter, Eli; Turner, Russell (2001-02-16). "The Sequence of the Human Genome". Science. 291 (5507): 1304–1351. Bibcode:2001Sci...291.1304V. doi:10.1126/science.1058040. ISSN 1095-9203. PMID 11181995.
- ^ Kang, Byoung-Cheorl; Nah, Gyoungju; Lee, Heung-Ryul; Han, Koeun; Purushotham, Preethi M.; Jo, Jinkwan (2017). "Development of a Genetic Map for Onion (Allium cepa L.) Using Reference-Free Genotyping-by-Sequencing and SNP Assays". Frontiers in Plant Science. 8: 1606. doi:10.3389/fpls.2017.01606. ISSN 1664-462X. PMC 5604068. PMID 28959273.
- ^ Smith, Jeramiah J.; Voss, S. Randal; Tsonis, Panagiotis A.; Timoshevskaya, Nataliya Y.; Timoshevskiy, Vladimir A.; Keinath, Melissa C. (2015-11-10). "Initial characterization of the large genome of the salamander Ambystoma mexicanum using shotgun and laser capture chromosome sequencing". Scientific Reports. 5: 16413. Bibcode:2015NatSR...516413K. doi:10.1038/srep16413. ISSN 2045-2322. PMC 4639759. PMID 26553646.
- ^ Venter, J. C.; Adams, M. D.; Myers, E. W.; Li, P. W.; Mural, R. J.; Sutton, G. G.; Smith, H. O.; Yandell, M.; Evans, C. A. (2001-02-16). "The sequence of the human genome". Science. 291 (5507): 1304–1351. Bibcode:2001Sci...291.1304V. doi:10.1126/science.1058040. ISSN 0036-8075. PMID 11181995.
- ^ Lander, E. S.; Linton, L. M.; Birren, B.; Nusbaum, C.; Zody, M. C.; Baldwin, J.; Devon, K.; Dewar, K.; Doyle, M. (2001-02-15). "Initial sequencing and analysis of the human genome" (PDF). Nature. 409 (6822): 860–921. Bibcode:2001Natur.409..860L. doi:10.1038/35057062. ISSN 0028-0836. PMID 11237011.
- ^ Consortium*, The C. elegans Sequencing (1998-12-11). "Genome Sequence of the Nematode C. elegans: A Platform for Investigating Biology". Science. 282 (5396): 2012–2018. Bibcode:1998Sci...282.2012.. doi:10.1126/science.282.5396.2012. ISSN 1095-9203. PMID 9851916.
- ^ Arabidopsis Genome Initiative (2000-12-14). "Analysis of the genome sequence of the flowering plant Arabidopsis thaliana". Nature. 408 (6814): 796–815. Bibcode:2000Natur.408..796T. doi:10.1038/35048692. ISSN 0028-0836. PMID 11130711.
- ^ Bennetzen, Jeffrey L.; Brown, James K. M.; Devos, Katrien M. (2002-07-01). "Genome Size Reduction through Illegitimate Recombination Counteracts Genome Expansion in Arabidopsis". Genome Research. 12 (7): 1075–1079. doi:10.1101/gr.132102. ISSN 1549-5469. PMC 186626. PMID 12097344.
- ^ Kurland, C. G.; Canbäck, B.; Berg, O. G. (December 2007). "The origins of modern proteomes". Biochimie. 89 (12): 1454–1463. doi:10.1016/j.biochi.2007.09.004. ISSN 0300-9084. PMID 17949885.
- ^ Caetano-Anollés, Gustavo; Caetano-Anollés, Derek (July 2003). "An evolutionarily structured universe of protein architecture". Genome Research. 13 (7): 1563–1571. doi:10.1101/gr.1161903. ISSN 1088-9051. PMC 403752. PMID 12840035.
- ^ Glansdorff, Nicolas; Xu, Ying; Labedan, Bernard (2008-07-09). "The last universal common ancestor: emergence, constitution and genetic legacy of an elusive forerunner". Biology Direct. 3: 29. doi:10.1186/1745-6150-3-29. ISSN 1745-6150. PMC 2478661. PMID 18613974.
- ^ Kurland, C. G.; Collins, L. J.; Penny, D. (2006-05-19). "Genomics and the irreducible nature of eukaryote cells". Science. 312 (5776): 1011–1014. Bibcode:2006Sci...312.1011K. doi:10.1126/science.1121674. ISSN 1095-9203. PMID 16709776. S2CID 30768101.
- ^ Collins, Lesley; Penny, David (April 2005). "Complex spliceosomal organization ancestral to extant eukaryotes". Molecular Biology and Evolution. 22 (4): 1053–1066. doi:10.1093/molbev/msi091. ISSN 0737-4038. PMID 15659557.
- ^ Penny, David; Collins, Lesley J.; Daly, Toni K.; Cox, Simon J. (December 2014). "The relative ages of eukaryotes and akaryotes". Journal of Molecular Evolution. 79 (5–6): 228–239. Bibcode:2014JMolE..79..228P. doi:10.1007/s00239-014-9643-y. ISSN 1432-1432. PMID 25179144. S2CID 17512331.
- ^ Fuerst, John A.; Sagulenko, Evgeny (2012-05-04). "Keys to Eukaryality: Planctomycetes and Ancestral Evolution of Cellular Complexity". Frontiers in Microbiology. 3: 167. doi:10.3389/fmicb.2012.00167. ISSN 1664-302X. PMC 3343278. PMID 22586422.
- ^ Collins, Lesley; Penny, David (April 2005). "Complex spliceosomal organization ancestral to extant eukaryotes". Molecular Biology and Evolution. 22 (4): 1053–1066. doi:10.1093/molbev/msi091. ISSN 0737-4038. PMID 15659557. [verification needed]
- ^ a b c d Shapiro, M. B.; Senapathy, P. (1987-09-11). "RNA splice junctions of different classes of eukaryotes: sequence statistics and functional implications in gene expression". Nucleic Acids Research. 15 (17): 7155–7174. doi:10.1093/nar/15.17.7155. ISSN 0305-1048. PMC 306199. PMID 3658675.
- ^ a b c d Senapathy, P.; Shapiro, M. B.; Harris, N. L. (1990). [16] Splice junctions, branch point sites, and exons: Sequence statistics, identification, and applications to genome project. Methods in Enzymology. Vol. 183. pp. 252–278. doi:10.1016/0076-6879(90)83018-5. ISBN 9780121820848. ISSN 0076-6879. PMID 2314278.
- ^ "National Institutes of Health (NIH) — All of Us". allofus.nih.gov. Retrieved 2019-01-02.
- ^ a b Penny, David; Collins, Lesley (2005-04-01). "Complex Spliceosomal Organization Ancestral to Extant Eukaryotes". Molecular Biology and Evolution. 22 (4): 1053–1066. doi:10.1093/molbev/msi091. ISSN 0737-4038. PMID 15659557.
- ^ Caetano-Anollés, Derek; Caetano-Anollés, Gustavo (2003-07-01). "An Evolutionarily Structured Universe of Protein Architecture". Genome Research. 13 (7): 1563–1571. doi:10.1101/gr.1161903. ISSN 1549-5469. PMC 403752. PMID 12840035.
- ^ Glansdorff, Nicolas; Xu, Ying; Labedan, Bernard (2008-07-09). "The Last Universal Common Ancestor: emergence, constitution and genetic legacy of an elusive forerunner". Biology Direct. 3 (1): 29. doi:10.1186/1745-6150-3-29. ISSN 1745-6150. PMC 2478661. PMID 18613974.
- ^ Kurland, C.G.; Canbäck, B.; Berg, O.G. (2007-12-01). "The origins of modern proteomes". Biochimie. 89 (12): 1454–1463. doi:10.1016/j.biochi.2007.09.004. ISSN 0300-9084. PMID 17949885.
- ^ Penny, D.; Collins, L. J.; Kurland, C. G. (2006-05-19). "Genomics and the Irreducible Nature of Eukaryote Cells". Science. 312 (5776): 1011–1014. Bibcode:2006Sci...312.1011K. doi:10.1126/science.1121674. ISSN 1095-9203. PMID 16709776. S2CID 30768101.
- ^ Poole, A. M.; Jeffares, D. C.; Penny, D. (January 1998). "The path from the RNA world". Journal of Molecular Evolution. 46 (1): 1–17. Bibcode:1998JMolE..46....1P. doi:10.1007/PL00006275. ISSN 0022-2844. PMID 9419221. S2CID 17968659.
- ^ Forterre, Patrick; Philippe, Hervé (1999). "Where is the root of the universal tree of life?". BioEssays. 21 (10): 871–879. doi:10.1002/(SICI)1521-1878(199910)21:10<871::AID-BIES10>3.0.CO;2-Q. ISSN 1521-1878. PMID 10497338.
- ^ Cox, Simon J.; Daly, Toni K.; Collins, Lesley J.; Penny, David (2014-12-01). "The Relative Ages of Eukaryotes and Akaryotes". Journal of Molecular Evolution. 79 (5–6): 228–239. Bibcode:2014JMolE..79..228P. doi:10.1007/s00239-014-9643-y. ISSN 1432-1432. PMID 25179144. S2CID 17512331.
- ^ Sagulenko, Evgeny; Fuerst, John Arlington (2012). "Keys to eukaryality: planctomycetes and ancestral evolution of cellular complexity". Frontiers in Microbiology. 3: 167. doi:10.3389/fmicb.2012.00167. ISSN 1664-302X. PMC 3343278. PMID 22586422.
- ^ a b Gilbert, Walter; Roy, Scott W. (2005-02-08). "Complex early genes". Proceedings of the National Academy of Sciences. 102 (6): 1986–1991. Bibcode:2005PNAS..102.1986R. doi:10.1073/pnas.0408355101. ISSN 1091-6490. PMC 548548. PMID 15687506.
- ^ "Introduction to the Basal Eukaryotes". ucmp.berkeley.edu. Retrieved 2021-08-01.
- ^ Gilbert, Walter; Roy, Scott William (March 2006). "The evolution of spliceosomal introns: patterns, puzzles and progress". Nature Reviews Genetics. 7 (3): 211–221. doi:10.1038/nrg1807. ISSN 1471-0064. PMID 16485020. S2CID 33672491.
- ^ Rogozin, Igor B.; Sverdlov, Alexander V.; Babenko, Vladimir N.; Koonin, Eugene V. (June 2005). "Analysis of evolution of exon-intron structure of eukaryotic genes". Briefings in Bioinformatics. 6 (2): 118–134. doi:10.1093/bib/6.2.118. ISSN 1467-5463. PMID 15975222.
- ^ Sullivan, James C.; Reitzel, Adam M.; Finnerty, John R. (2006). "A high percentage of introns in human genes were present early in animal evolution: Evidence from the basal metazoan Nematostella vectensis". Genome Informatics. International Conference on Genome Informatics. 17 (1): 219–229. ISSN 0919-9454. PMID 17503371.
- ^ Koonin, Eugene V.; Rogozin, Igor B.; Csuros, Miklos (2011-09-15). "A Detailed History of Intron-rich Eukaryotic Ancestors Inferred from a Global Survey of 100 Complete Genomes". PLOS Computational Biology. 7 (9): e1002150. Bibcode:2011PLSCB...7E2150C. doi:10.1371/journal.pcbi.1002150. ISSN 1553-7358. PMC 3174169. PMID 21935348.
