Dictyostelid
| Dictyostelid | |
|---|---|

| |
| Dictyostelium discoideum | |
| Scientific classification | |
| Domain: | Eukaryota |
| Phylum: | Amoebozoa |
| Subphylum: | Conosa |
| Infraphylum: | Eumycetozoa |
| Class: | Dictyostelia Lister 1909, emend. Olive 1970 |
| Orders | |
The dictyostelids (Dictyostelia/Dictyostelea, ICZN, Dictyosteliomycetes, ICBN) or cellular slime molds are a group of slime molds or social amoebae.
Multicellular behavior
[edit]
When food (normally bacteria) is readily available dictyostelids behave as individual amoebae, which feed and divide normally. However, when the food supply is exhausted, they aggregate to form a multicellular assembly, called a pseudoplasmodium, grex, or slug (not to be confused with the gastropod mollusc called a slug). The grex has a definite anterior and posterior, responds to light and temperature gradients, and has the ability to migrate. Under the correct circumstances the grex matures forming a sorocarp (fruiting body) with a stalk supporting one or more sori (balls of spores). These spores are inactive cells protected by resistant cell walls, and become new amoebae once food is available.
In Acytostelium, the sorocarp is supported by a stalk composed of cellulose, but in other dictyostelids the stalk is composed of cells, sometimes taking up the majority of the original amoebae. With a few exceptions, these cells die during stalk formation, and there is a definite correspondence between parts of the grex and parts of the fruiting body. Aggregation of amoebae generally takes place in converging streams. The amoebae move using filose pseudopods, and are attracted to chemicals produced by other amoebae. In Dictyostelium discoideum, aggregation is signalled by cAMP, but others use different chemicals. In the species Dictyostelium purpureum, the grouping is by kinship, not just by proximity.
Uses as model organism
[edit]Dictyostelium has been used as a model organism in molecular biology and genetics, and is studied as an example of cell communication, differentiation, and programmed cell death. It is also an interesting example of the evolution of cooperation and cheating.[1][2][3] A large body of research data concerning D. discoideum is available on-line at DictyBase.

Mechanism of aggregation in Dictyostelium discoideum
[edit]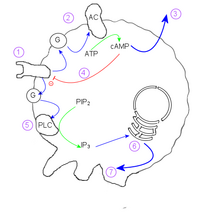
The mechanism behind the aggregation of the amoebae relies on cyclic adenosine monophosphate (cAMP) as a signal molecule. One cell, the founder of the colony, begins to secrete cAMP in response to stress. Others detect this signal, and respond in two ways:
- The amoeba moves towards the signal.
- The amoeba secretes more cAMP to boost the signal.
The effect of this is to relay the signal throughout the nearby population of amoebae and cause inward movement to the area of highest cAMP concentration.
Within an individual cell, the mechanism is as follows:
- cAMP reception at the cell membrane activates a G-protein
- G protein stimulates Adenylate cyclase
- cAMP diffuses out of cell into medium
- Internal cAMP inactivates the external cAMP receptor.
- A different g-protein stimulates Phospholipase C
- IP3 induces calcium ion release
- Calcium ions act on the cytoskeleton to induce the extension of pseudopodia.
Because the internal cAMP concentration inactivates the receptor for external cAMP, an individual cell shows oscillatory behaviour. This behaviour produces beautiful spirals seen in converging colonies and is reminiscent of the Belousov–Zhabotinsky reaction and two-dimensional cyclic cellular automata.
Genome
[edit]The entire genome of Dictyostelium discoideum was published in Nature in 2005 by geneticist Ludwig Eichinger and coworkers.[4] The haploid genome contains approximately 12,500 genes on 6 chromosomes. For comparison, the diploid human genome has 20,000-25,000 genes (represented twice) on 23 chromosome pairs. There is a high level of the nucleotides adenosine and thymidine (~77%) leading to a codon usage that favors more adenosines and thymidines in the third position. Tandem repeats of trinucleotides are abundant in Dictyostelium, which in humans cause Trinucleotide repeat disorders.
Sexual reproduction
[edit]Sexual development can occur when amoeboid cells are starved for their bacterial food supply and dark humid conditions are present.[5] Both heterothallic and homothallic strains of Dictostelium can undergo mating. Heterothallic sexual development has been most extensively studied in D. discoideum, and homothallic sexual development has been most well studied in D. mucoroides.[6] Heterothallic matings are initiated by fusion of haploid cells (gametes) from two strains of opposite mating type. This contrasts with homothallic strains that appear to express both mating types.[7]
Mating is initiated by gametogenesis that produces small, motile gametes that fuse to form a small binucleate cell. The volume of the binucleate cell then increases to produce a giant binuclear cell. As growth proceeds, the nuclei swell, and then fuse forming a true diploid zygote giant cell. As this is occurring, amoebae have been undergoing cAMP-induced chemotaxis towards the giant cell surface. This forms a cellular aggregate and at the center of the aggregate the zygote giant cell ingests the surrounding amoebae. Phagocytosis is followed by digestion of the ingested amoebae. Next the zygote forms a macrocyst characterized by a surrounding extracellular cellulose sheath. After the macrocyst is formed it ordinarily remains dormant for a period before germination can occur.[8] Within the macrocyst the diploid zygote undergoes meiosis followed by successive mitotic divisions. When the macrocyst germinates it releases many haploid amoeboid cells.
Taxonomy
[edit]Classification History
[edit]The Dictyostelium phylogeny tree has undergone multiple changes in the past decades. The first dictyostelid to be described was Dictyostelium mucoroides in 1869 by Osker Brefeld, and the original discovery of Dictyostelium discoideum occurred in 1935,[9] with further discoveries from Kenneth Raper, followed by global efforts led by James Cavender and collaborators. Dictyostelium discoideum was initially classified under ‘lower fungi,’ but the classification has since shifted the classification to under the Amoebozoa phylum where it is currently.[4]
The groupings within the dictyostelid phylogeny tree has undergone frequent reordering due to availability new evidence. Most currently accepted phylogenies of dictyostelids utilize genome sequencing and small subunit ribosomal DNA (ssu-rDNA). The dictyostelids can be further subdivided into four groups. In particular, group 4 contains the Dictyostelium discoideum species and differs from the other groups with its usage of cAMP as an attractant emitted during aggregation.[10]
Evolution
[edit]Fossil calibrations indicate that dictyostelids class originally diverged into two major branches approximately 0.52 billion years ago. Current theories speculate that dictyostelid stalk and spore formation originally evolved as an adaption to global glacial formations. Further subdivision of dictyostelids species likely arose as most glacial formations melted. Most species in major groups 1, 2, and 3 display an encystment ability that allows individuals to survive low temperatures, but spores have shown an increased ability in resisting lower temperatures. Group 4 differs from other major groups in a general lacking ability to encyst, but its spores have shown better resistances against lower temperatures relative to spores from other groups.[11]
The internal phylogeny of the Dictyostelids is shown in the cladogram. [12]
| Dictyosteliida | |
Taxonomy
[edit]Class Dictyostelia Lister 1909 em. Olive 1970[13]
- Genus ?Calospeira Arnaud 1949
- Genus ?Coenonia van Tieghem 1884 non Vandamme et al. 1999
- Genus ?Synstelium Baldauf, Sheikh & Thulin 2017
- Order Actyosteliales Baldauf, Sheikh & Thulin 2017
- Family Cavenderiaceae Baldauf, Sheikh & Thulin 2017
- Genus Cavenderia Baldauf, Sheikh & Thulin 2017
- Family Acytosteliidae Raper ex Raper & Quinlan 1958
- Genus Acytostelium Raper 1956
- Genus Heterostelium Baldauf, Sheikh & Thulin 2017
- Genus Rostrostelium Baldauf, Sheikh & Thulin 2017
- Family Cavenderiaceae Baldauf, Sheikh & Thulin 2017
- Order Dictyosteliales Olive ex Kirk, Cannon & David 2001
- Genus ?Coremiostelium Baldauf, Sheikh & Thulin 2017
- Genus ?Heliomycopsis Arnaud 1949
- Genus ?Pygmomyces Arnaud 1949
- Genus †?Myxomitodes Bengtson et al. 2007
- Genus ?Rhabdocystis Arnaud 1949
- Family Dictyosteliidae Rostafinski 1875 ex Cooke 1877
- Dictyostelium Brefeld 1870
- Polysphondylium Brefeld 1884
- Family Raperosteliaceae Baldauf, Sheikh & Thulin 2017
- Genus Hagiwaraea Baldauf, Sheikh & Thulin 2017
- Genus Raperostelium Baldauf, Sheikh & Thulin 2017
- Genus Speleostelium Baldauf, Sheikh & Thulin 2017
- Genus Tieghemostelium Baldauf, Sheikh & Thulin 2017
Model host organism for Legionella
[edit]Dictyostelium shares many molecular features with macrophages, the human host of Legionella. The cytoskeletal composition of D. discoideum is similar to that of mammalian cells as are the processes driven by these components, such as phagocytosis, membrane trafficking, endocytic transit and vesicle sorting. Like leukocytes, D. discoideum possess chemotactic capacity. Hence, D. discoideum represents a suitable model system to ascertain the influence of a variety of host cell factors during Legionella infections.[14]
References
[edit]- ^ Strassman JE, Zhu Y, and Queller DC. (2000) Altruism and social cheating in the social amoeba Dictyostelium discoideum. Nature
- ^ Dao DN, Kessin RH, and Ennis HL (2000). Developmental cheating and the evolutionary biology of Dictyostelium and Myxococcus. Microbiology
- ^ Brännsröm Å and Dieckmann U (2005). Evolutionary dynamics of altruism and cheating among social amoebas. Proceedings of the Royal Society of London B.
- ^ a b Eichinger, L.; Pachebat, J.A.; Glöckner, G.; Rajandream, M.A.; Sucgang, R.; Berriman, M.; Song, J.; Olsen, R.; Szafranski, K.; Xu, Q.; et al. (2005). "The genome of the social amoeba Dictyostelium discoideum". Nature. 435 (7038): 43–57. Bibcode:2005Natur.435...43E. doi:10.1038/nature03481. PMC 1352341. PMID 15875012.
- ^ Flowers JM, Li SI, Stathos A, Saxer G, Ostrowski EA, Queller DC, Strassmann JE, Purugganan MD (July 2010). "Variation, sex, and social cooperation: molecular population genetics of the social amoeba Dictyostelium discoideum". PLOS Genet. 6 (7): e1001013. doi:10.1371/journal.pgen.1001013. PMC 2895654. PMID 20617172.
- ^ O'Day DH, Keszei A (May 2012). "Signalling and sex in the social amoebozoans". Biol Rev Camb Philos Soc. 87 (2): 313–29. doi:10.1111/j.1469-185X.2011.00200.x. PMID 21929567. S2CID 205599638.
- ^ Robson GE, Williams KL (April 1980). "The mating system of the cellular slime mould Dictyostelium discoideum". Curr. Genet. 1 (3): 229–32. doi:10.1007/BF00390948. PMID 24189663. S2CID 23172357.
- ^ Nickerson AW, Raper KB. Macrocysts in the life cycle of Dictyostelliaceae II. Germination of the macrocysts. Am. J. Bot. 1973 60(3): 247–254.
- ^ Brefeld, O (1869). "Ein neuer Organismus und der Verwandschaft der Myxomyceten". Abh Seckenberg Naturforsch Ges. 7: 85–107.
- ^ Schilde, Christina; Lawal, Hajara M.; Kin, Koryu; Shibano-Hayakawa, Ikumi; Inouye, Kei; Schaap, Pauline (2019). "A well supported multi gene phylogeny of 52 dictyostelia". Molecular Phylogenetics and Evolution. 134: 66–73. doi:10.1016/j.ympev.2019.01.017. ISSN 1055-7903. PMC 6430600. PMID 30711536.
- ^ Lawal, Hajara M.; Schilde, Christina; Kin, Koryu; Brown, Matthew W.; James, John; Prescott, Alan R.; Schaap, Pauline (2020-05-29). "Cold climate adaptation is a plausible cause for evolution of multicellular sporulation in Dictyostelia". Scientific Reports. 10 (1): 8797. doi:10.1038/s41598-020-65709-3. ISSN 2045-2322. PMC 7260361. PMID 32472019.
- ^ Sheikh, Sanea; Thulin, Mats; Cavender, James C; Escalante, Ricardo; Kawakami, Shin-ichi; Lado, Carlos; Landolt, John; Nanjundiah, Vidyanand; Queller, David; Strassmann, Joan; Spiegel, Frederick W.; Stephenson, Steve; Vadell, Eduardo M; Baldauf, Sandra (2017-11-24). "A New Classification of the Dictyostelids". Protists. 169 (1): 1–28. doi:10.1016/j.protis.2017.11.001. PMID 29367151.
- ^ Wijayawardene, Nalin; Hyde, Kevin; Al-Ani, LKT; Dolatabadi, S; Stadler, Marc; Haelewaters, Danny; et al. (2020). "Outline of Fungi and fungus-like taxa". Mycosphere. 11: 1060–1456. doi:10.5943/mycosphere/11/1/8. hdl:10481/61998.
- ^ Bruhn; et al. (2008). "Dictyostelium, a Tractable Model Host Organism for Legionella". Legionella: Molecular Microbiology. Caister Academic Press. ISBN 978-1-904455-26-4.
External links
[edit]- Dictyostelium (2007)
- Low Society (2004)
- dictyBase Online Informatics Resource for Dictyostelium
- dictyBase wiki official wiki site for dictyBase
- Dictyostelium discoideum Genome Project Archived 2019-12-06 at the Wayback Machine
- Dictyostelium discoideum description, life cycle
