Nucleic acid structure


Nucleic acid structure refers to the structure of nucleic acids such as DNA and RNA. Chemically speaking, DNA and RNA are very similar. Nucleic acid structure is often divided into four different levels: primary, secondary, tertiary, and quaternary.
Primary structure
[edit]
Primary structure consists of a linear sequence of nucleotides that are linked together by phosphodiester bond. It is this linear sequence of nucleotides that make up the primary structure of DNA or RNA. Nucleotides consist of 3 components:
- Nitrogenous base
- 5-carbon sugar which is called deoxyribose (found in DNA) and ribose (found in RNA).
- One or more phosphate groups.[1]
The nitrogen bases adenine and guanine are purine in structure and form a glycosidic bond between their 9 nitrogen and the 1' -OH group of the deoxyribose. Cytosine, thymine, and uracil are pyrimidines, hence the glycosidic bonds form between their 1 nitrogen and the 1' -OH of the deoxyribose. For both the purine and pyrimidine bases, the phosphate group forms a bond with the deoxyribose sugar through an ester bond between one of its negatively charged oxygen groups and the 5' -OH of the sugar.[2] The polarity in DNA and RNA is derived from the oxygen and nitrogen atoms in the backbone. Nucleic acids are formed when nucleotides come together through phosphodiester linkages between the 5' and 3' carbon atoms.[3] A nucleic acid sequence is the order of nucleotides within a DNA (GACT) or RNA (GACU) molecule that is determined by a series of letters. Sequences are presented from the 5' to 3' end and determine the covalent structure of the entire molecule. Sequences can be complementary to another sequence in that the base on each position is complementary as well as in the reverse order. An example of a complementary sequence to AGCT is TCGA. DNA is double-stranded containing both a sense strand and an antisense strand. Therefore, the complementary sequence will be to the sense strand.[4]
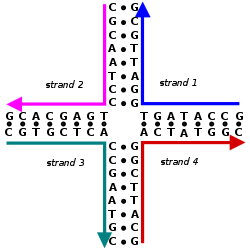
Complexes with alkali metal ions
[edit]There are three potential metal binding groups on nucleic acids: phosphate, sugar, and base moieties. Solid-state structure of complexes with alkali metal ions have been reviewed.[6]
Secondary structure
[edit]DNA
[edit]Secondary structure is the set of interactions between bases, i.e., which parts of strands are bound to each other. In DNA double helix, the two strands of DNA are held together by hydrogen bonds. The nucleotides on one strand base pairs with the nucleotide on the other strand. The secondary structure is responsible for the shape that the nucleic acid assumes. The bases in the DNA are classified as purines and pyrimidines. The purines are adenine and guanine. Purines consist of a double ring structure, a six-membered and a five-membered ring containing nitrogen. The pyrimidines are cytosine and thymine. It has a single ring structure, a six-membered ring containing nitrogen. A purine base always pairs with a pyrimidine base (guanine (G) pairs with cytosine (C) and adenine (A) pairs with thymine (T) or uracil (U)). DNA's secondary structure is predominantly determined by base-pairing of the two polynucleotide strands wrapped around each other to form a double helix. Although the two strands are aligned by hydrogen bonds in base pairs, the stronger forces holding the two strands together are stacking interactions between the bases. These stacking interactions are stabilized by Van der Waals forces and hydrophobic interactions, and show a large amount of local structural variability.[7] There are also two grooves in the double helix, which are called major groove and minor groove based on their relative size.
RNA
[edit]
The secondary structure of RNA consists of a single polynucleotide. Base pairing in RNA occurs when RNA folds between complementarity regions. Both single- and double-stranded regions are often found in RNA molecules.
The four basic elements in the secondary structure of RNA are:
- Helices
- Bulges
- Loops
- Junctions
The antiparallel strands form a helical shape.[3] Bulges and internal loops are formed by separation of the double helical tract on either one strand (bulge) or on both strands (internal loops) by unpaired nucleotides.
Stem-loop or hairpin loop is the most common element of RNA secondary structure.[8] Stem-loop is formed when the RNA chains fold back on themselves to form a double helical tract called the 'stem', the unpaired nucleotides forms single stranded region called the 'loop'. A tetraloop is a four-base pairs hairpin RNA structure. There are three common families of tetraloop in ribosomal RNA: UNCG, GNRA, and CUUG (N is one of the four nucleotides and R is a purine). UNCG is the most stable tetraloop.[9]
Pseudoknot is an RNA secondary structure first identified in turnip yellow mosaic virus.[10] It is minimally composed of two helical segments connected by single-stranded regions or loops. H-type fold pseudoknots are best characterized. In H-type fold, nucleotides in the hairpin-loop pair with the bases outside the hairpin stem forming second stem and loop. This causes formation of pseudoknots with two stems and two loops.[11] Pseudoknots are functional elements in RNA structure having diverse function and found in most classes of RNA.
Secondary structure of RNA can be predicted by experimental data on the secondary structure elements, helices, loops, and bulges. DotKnot-PW method is used for comparative pseudoknots prediction. The main points in the DotKnot-PW method is scoring the similarities found in stems, secondary elements and H-type pseudoknots.[12]
Tertiary structure
[edit]

Tertiary structure refers to the locations of the atoms in three-dimensional space, taking into consideration geometrical and steric constraints. It is a higher order than the secondary structure, in which large-scale folding in a linear polymer occurs and the entire chain is folded into a specific 3-dimensional shape. There are 4 areas in which the structural forms of DNA can differ.
- Handedness – right or left
- Length of the helix turn
- Number of base pairs per turn
- Difference in size between the major and minor grooves[3]
The tertiary arrangement of DNA's double helix in space includes B-DNA, A-DNA, and Z-DNA. Triple-stranded DNA structures have been demonstrated in repetitive polypurine:polypyrimidine Microsatellite sequences and Satellite DNA.
B-DNA is the most common form of DNA in vivo and is a more narrow, elongated helix than A-DNA. Its wide major groove makes it more accessible to proteins. On the other hand, it has a narrow minor groove. B-DNA's favored conformations occur at high water concentrations; the hydration of the minor groove appears to favor B-DNA. B-DNA base pairs are nearly perpendicular to the helix axis. The sugar pucker which determines the shape of the a-helix, whether the helix will exist in the A-form or in the B-form, occurs at the C2'-endo.[13]
A-DNA, is a form of the DNA duplex observed under dehydrating conditions. It is shorter and wider than B-DNA. RNA adopts this double helical form, and RNA-DNA duplexes are mostly A-form, but B-form RNA-DNA duplexes have been observed.[14] In localized single strand dinucleotide contexts, RNA can also adopt the B-form without pairing to DNA.[15] A-DNA has a deep, narrow major groove which does not make it easily accessible to proteins. On the other hand, its wide, shallow minor groove makes it accessible to proteins but with lower information content than the major groove. Its favored conformation is at low water concentrations. A-DNAs base pairs are tilted relative to the helix axis, and are displaced from the axis. The sugar pucker occurs at the C3'-endo and in RNA 2'-OH inhibits C2'-endo conformation.[13] Long considered little more than a laboratory artifice, A-DNA is now known to have several biological functions.
Z-DNA is a relatively rare left-handed double-helix. Given the proper sequence and superhelical tension, it can be formed in vivo but its function is unclear. It has a more narrow, more elongated helix than A or B. Z-DNA's major groove is not really a groove, and it has a narrow minor groove. The most favored conformation occurs when there are high salt concentrations. There are some base substitutions but they require an alternating purine-pyrimidine sequence. The N2-amino of G H-bonds to 5' PO, which explains the slow exchange of protons and the need for the G purine. Z-DNA base pairs are nearly perpendicular to the helix axis. Z-DNA does not contain single base-pairs but rather a GpC repeat with P-P distances varying for GpC and CpG. On the GpC stack there is good base overlap, whereas on the CpG stack there is less overlap. Z-DNA's zigzag backbone is due to the C sugar conformation compensating for G glycosidic bond conformation. The conformation of G is syn, C2'-endo; for C it is anti, C3'-endo.[13]
A linear DNA molecule having free ends can rotate, to adjust to changes of various dynamic processes in the cell, by changing how many times the two chains of its double helix twist around each other. Some DNA molecules are circular and are topologically constrained. More recently circular RNA was described as well to be a natural pervasive class of nucleic acids, expressed in many organisms (see CircRNA).
A covalently closed, circular DNA (also known as cccDNA) is topologically constrained as the number of times the chains coiled around one other cannot change. This cccDNA can be supercoiled, which is the tertiary structure of DNA. Supercoiling is characterized by the linking number, twist and writhe. The linking number (Lk) for circular DNA is defined as the number of times one strand would have to pass through the other strand to completely separate the two strands. The linking number for circular DNA can only be changed by breaking of a covalent bond in one of the two strands. Always an integer, the linking number of a cccDNA is the sum of two components: twists (Tw) and writhes (Wr).[16]
Twists are the number of times the two strands of DNA are twisted around each other. Writhes are number of times the DNA helix crosses over itself. DNA in cells is negatively supercoiled and has the tendency to unwind. Hence the separation of strands is easier in negatively supercoiled DNA than in relaxed DNA. The two components of supercoiled DNA are solenoid and plectonemic. The plectonemic supercoil is found in prokaryotes, while the solenoidal supercoiling is mostly seen in eukaryotes.
Quaternary structure
[edit]The quaternary structure of nucleic acids is similar to that of protein quaternary structure. Although some of the concepts are not exactly the same, the quaternary structure refers to a higher-level of organization of nucleic acids. Moreover, it refers to interactions of the nucleic acids with other molecules. The most commonly seen form of higher-level organization of nucleic acids is seen in the form of chromatin which leads to its interactions with the small proteins histones. Also, the quaternary structure refers to the interactions between separate RNA units in the ribosome or spliceosome.[17]
See also
[edit]- Biomolecular structure
- Crosslinking of DNA
- DNA nanotechnology
- DNA supercoil
- Gene structure
- Non-helical models of DNA structure
- Nucleic acid design
- Nucleic acid double helix
- Nucleic acid structure determination (experimental)
- Nucleic acid structure prediction (computational)
- Nucleic acid thermodynamics
- Protein structure
- Satellite DNA
- Triple-stranded DNA
References
[edit]- ^ Krieger M, Scott MP, Matsudaira PT, Lodish HF, Darnell JE, Lawrence Z, Kaiser C, Berk A (2004). "Section 4.1: Structure of Nucleic Acids". Molecular cell biology. New York: W.H. Freeman and CO. ISBN 978-0-7167-4366-8.
- ^ "Structure of Nucleic Acids". SparkNotes.
- ^ a b c Anthony-Cahill SJ, Mathews CK, van Holde KE, Appling DR (2012). Biochemistry (4th ed.). Englewood Cliffs, N.J: Prentice Hall. ISBN 978-0-13-800464-4.
- ^ Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Wlater P (2002). Molecular Biology of the Cell (4th ed.). New York NY: Garland Science. ISBN 978-0-8153-3218-3.
- ^ Mao C (December 2004). "The emergence of complexity: lessons from DNA". PLOS Biology. 2 (12): e431. doi:10.1371/journal.pbio.0020431. PMC 535573. PMID 15597116.
- ^ Katsuyuki, Aoki; Kazutaka, Murayama; Hu, Ning-Hai (2016). "Solid State Structures of Alkali Metal Ion Complexes Formed by Low-Molecular-Weight Ligands of Biological Relevance". In Astrid, Sigel; Helmut, Sigel; Roland K.O., Sigel (eds.). The Alkali Metal Ions: Their Role for Life. Metal Ions in Life Sciences. Vol. 16. Springer. pp. 43–66. doi:10.1007/978-3-319-21756-7_3. ISBN 978-3-319-21755-0. PMID 26860299.
- ^ Sedova A, Banavali NK (2017). "Geometric Patterns for Neighboring Bases Near the Stacked State in Nucleic Acid Strands". Biochemistry. 56 (10): 1426–1443. doi:10.1021/acs.biochem.6b01101. PMID 28187685.
- ^ Tinoco I, Bustamante C (October 1999). "How RNA folds". Journal of Molecular Biology. 293 (2): 271–81. doi:10.1006/jmbi.1999.3001. PMID 10550208.
- ^ Hollyfield JG, Besharse JC, Rayborn ME (December 1976). "The effect of light on the quantity of phagosomes in the pigment epithelium". Experimental Eye Research. 23 (6): 623–35. doi:10.1016/0014-4835(76)90221-9. PMID 1087245.
- ^ Rietveld K, Van Poelgeest R, Pleij CW, Van Boom JH, Bosch L (March 1982). "The tRNA-like structure at the 3' terminus of turnip yellow mosaic virus RNA. Differences and similarities with canonical tRNA". Nucleic Acids Research. 10 (6): 1929–46. doi:10.1093/nar/10.6.1929. PMC 320581. PMID 7079175.
- ^ Staple DW, Butcher SE (June 2005). "Pseudoknots: RNA structures with diverse functions". PLOS Biology. 3 (6): e213. doi:10.1371/journal.pbio.0030213. PMC 1149493. PMID 15941360.
- ^ Sperschneider J, Datta A, Wise MJ (December 2012). "Predicting pseudoknotted structures across two RNA sequences". Bioinformatics. 28 (23): 3058–65. doi:10.1093/bioinformatics/bts575. PMC 3516145. PMID 23044552.
- ^ a b c Dickerson RE, Drew HR, Conner BN, Wing RM, Fratini AV, Kopka ML (April 1982). "The anatomy of A-, B-, and Z-DNA". Science. 216 (4545): 475–85. Bibcode:1982Sci...216..475D. doi:10.1126/science.7071593. PMID 7071593.
- ^ Chen X; Ramakrishnan B; Sundaralingam M (1995). "Crystal structures of B-form DNA-RNA chimers complexed with distamycin". Nature Structural Biology. 2 (9): 733–735. doi:10.1038/nsb0995-733. PMID 7552741. S2CID 6886088.
- ^ Sedova A, Banavali NK (2016). "RNA approaches the B-form in stacked single strand dinucleotide contexts". Biopolymers. 105 (2): 65–82. doi:10.1002/bip.22750. PMID 26443416. S2CID 35949700.
- ^ Mirkin SM (2001). "DNA Topology: Fundamentals". eLS. doi:10.1038/npg.els.0001038. ISBN 978-0470016176.
{{cite book}}:|journal=ignored (help) - ^ "Structural Biochemistry/Nucleic Acid/DNA/DNA structure". Retrieved 11 December 2012.

